Investigation of novel SARS-CoV-2 variant
Variant of Concern 2021/03
Technical briefing 7
This briefing provides an update on previous briefings up to 13 February 2021
Summary
• There are 4 variants of concern and 5 variants under investigation (Table 1)
• A slightly increased risk of hospitalisation has been now been detected for VOC
202012/01 (B.1.1.7), supporting previous findings on fatality.
• The first VOC 202101/02 (P.1) cases have been detected in England. The VOC
202101/02 (P.1) risk assessment indicates that it is plausible that there is some
degree of either immune escape or increased transmissibility, or both. This is
based on laboratory data and modelling; the magnitude and clinical significance
of these effects have yet to be determined.
• VOCs (excluding VOC 202012/01 B.1.1.7) and VUIs remain low at less than 1%
of samples with a sequence result available.
| England genomic cases 10 February 2021 | ||||
| Variant | Pangolin lineage | confirmed | probable | total confirmed and probable |
| VOC 202012/01 | B1.1.7 | 98,422 | 10,671 | 109,093 |
| VOC 202102/02 | B.1.1.7 with E484K cluster | 34 | 0 | 34 |
| VOC 202012/02 | B.1.351 | 190 | 76 | 266 |
| VOC 202101/02 | P1 | 5 | 2 | 7 |
| VUI 202102/01 | A.23.1 with E484K | 59 | 19 | 78 |
| VUI 202102/03 | B.1.525 | 79 | 28 | 107 |
| VUI 202102/04 | B.1.1.318 | 18 | 0 | 18 |
| VUI 202103/01 | B.1.324.1 with E484K | 2 | 0 | 2 |
Case numbers on variants of concern (VOC) and variants under investigation (VUI) are
updated online. Note table excludes variant cases not linked to a known COVID-19 case,
which may represent duplicate results or non-England residents.
Variants in Monitoring
Variants in monitoring are those which have been identified through horizon scanning, do not
have sufficient clear signals of concern to escalate further, but for which case numbers and
available data are being monitored regularly. These variants are in the monitoring category
• B.1.429 first detected in California (10/329,734 UK sequences as of 10 March 2021)
• B.1.1.7 with S494P first detected in the UK (761/329,734 UK sequences as of 10 March 2021)
• A.27 first detected in Mayotte (4/329,734 UK sequences as of 10 March 2021)
• B.1.526 first detected in New York (2/329,734 UK sequences as of 10 March 2021)
Variants in Monitoring
Repository of human and machine readable genomic case definitions
A repository containing the up-to-date genomic definitions for all VOC and VUI as curated by
Public Health England was created 5 March 2021. The repository can be accessed on
GitHub. They are provided in order to facilitate standardised VOC and VUI calling across
sequencing sites and bioinformatics pipelines and are the same definitions used internally at
Public Health England. Definition files are provided in YAML format so are compatible with a
range of computational platforms. The repository will be regularly updated.
Variant risk assessment includes the following confidence grading categorisations and utilises the framework in Table 2.
• Low: Little or poor-quality evidence, uncertainty or conflicting views amongst experts, no experience with previous similar incidents.
• Moderate: Adequate quality evidence, including consistent results published only in grey literature, reliable source(s), assumptions made on analogy and agreement between experts or opinion of at least 2 trusted experts.
• High: Good quality evidence, multiple reliable sources, verified, expert opinion concurs, experience of previous similar incidents.
| Indicator | Risk assessment framework | |||
| Zoonotic emergence and transmission to humans | Animal reservoir identified but no evidence of transmission from animals to humans | Sporadic transmission from animals to humans | Frequent transmission from animals to humans | |
| Transmissibility between humans | No demonstrated person to person transmission | Limited human case clusters | Established human to human transmission, which appears similar to wild type virus | Transmissibility appears greater than the wild type virus |
| Infection severity | Evidence of less severe clinical picture or lower infection fatality than from wild type SARS-CoV-2 infections | Similar clinical picture and infection fatality to wild type SARS-CoV-2 infections OR ex perimental animal data suggesting potential for increased disease severity in humans | More severe clinical picture or higher infection fatality than from wild type SARS-CoV-2 infections (limited to specific risk groups) | More severe clinical picture or higher infection fatality than from wild type SARSCoV-2 infections |
| Susceptibility and immunity – natural infection | Evidence of no antigenic difference from other circulating wild type virus | Structural data suggesting antigenic difference from other circulating wild type virus | Experimental evidence of functional evasion of naturally acquired immunity | Evidence of frequent infection in humans with known prior infection with earlier virus variant. |
| Vaccines | Evidence of no structural or antigenic difference in vaccine targets | Structural data suggesting difference in vaccine target epitopes | Experimental evidence of functional evasion of vaccine derived immunity | Evidence of frequent vaccine failure or decreased effectiveness in humans |
| Drugs and therapeutics | Evidence of no structural or antigenic difference in therapeutic targets | Structural data suggesting difference in therapeutic target epitopes | Experimental evidence of reduced drug susceptibility | Evidence of frequent drug or therapeutic failure or decreased effectiveness in humans |
VOC 202012/01 (B.1.1.7)
This variant was designated VUI 202012/01 (B.1.1.7) on detection and on review redesignated
as VOC 202012/01 (B.1.1.7) on 18 December 2020.
Genomic profile
Lineage defining mutations are shown in technical briefing 6. In addition, VOC 202012/01 has
acquired other mutations in some cases. Mutation counts in the UK dataset are shown in
Table 3.
| VOC 202012/01 (B.1.1.7) Spike variants | ||||
| Amino acid change | Total number of instances in VOC 202012/01 (B.1.1.7) (UK data) to 10 March 2021 | 11 December 2020 to 10 January 2021 | 11 January 2021 to 10 February 2021 | 11 February 2021 to 10 March 2021 |
| L18F | 3720(2.7%) | 314(0.7%) | 1431(2.1%) | 549(1.7%) |
| Q677H | 851(0.6%) | 92(0.2%) | 366(0.5%) | 348(1.1%) |
| S494P | 761(0.6%) | 208(0.4%) | 339(0.5%) | 88(0.3%) |
| G142V | 131(0.1%) | 8(0%) | 25(0%) | 84(0.3%) |
| S680F | 103(0.1%) | 13(0%) | 36(0.1%) | 42(0.1%) |
| Y144F | 354(0.3%) | 120(0.3%) | 164(0.2%) | 37(0.1%) |
| T678I | 144(0.1%) | 25(0.1%) | 54(0.1%) | 30(0.1%) |
| H146Y | 70(0.1%) | 7(0%) | 23(0%) | 27(0.1%) |
| F490S | 109(0.1%) | 10(0%) | 64(0.1%) | 24(0.1%) |
| A475A | 121(0.1%) | 11(0%) | 64(0.1%) | 23(0.1%) |
| E484K | 65(0%) | 10(0%) | 37(0.1%) | 13(0%) |
| Q493K | 14(0%) | 0(0%) | 1(0%) | 11(0%) |
| A684V | 51(0%) | 8(0%) | 23(0%) | 10(0%) |
| Total VOC 202012/01 (B.1.1.7) 135,310 | ||||
Epidemiological profile
Lineage B.1.1.7 is dispersed across the UK. Confirmed cases are those identified by whole
genome sequencing. As of 10 March 2021, there were 109,093 confirmed and probable cases
of VOC 202012/01 (B.1.1.7) in England (98,422 confirmed and 10,671 probable). The use of S
gene target failure in the Taqpath assay as a good proxy for VOC 202012/01 (B.1.1.7) cases
has been described in prior technical briefings. This continues to be supported by current data
(Appendix 1). In samples tested with this assay in the Lighthouse Laboratories, samples with
SGTF have predominated since mid December 2020, reaching 99% of cases in the week
starting 3 March 2021. Proportions in all regions are >97% (Appendix 1). Confirmed and
probable cases by specimen date are shown in Figure 1 and age-sex pyramid in Figure 2.
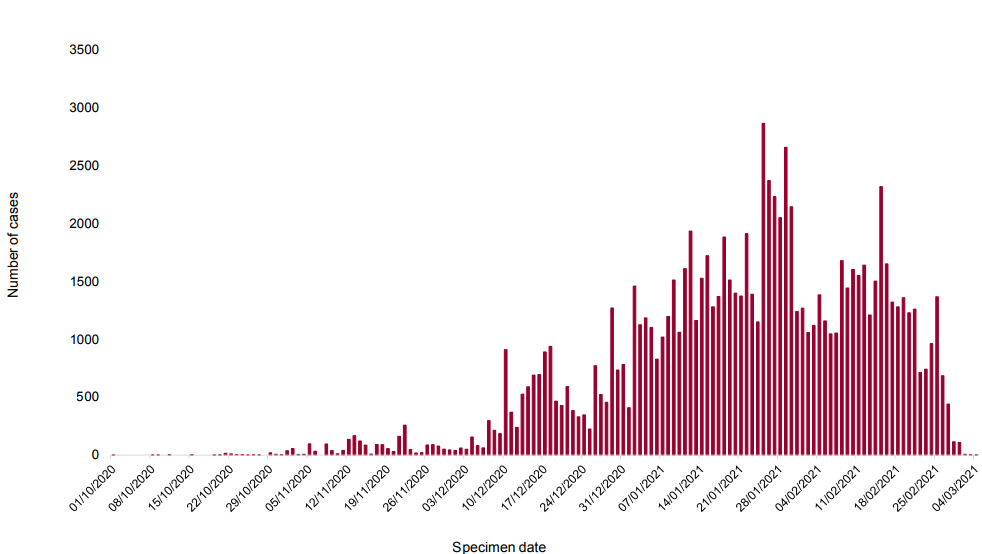
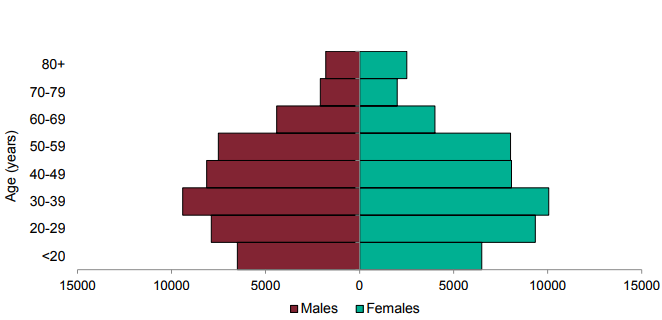
Figure 2. Age-sex pyramid of VOC 202012/01 (B.1.1.7) cases, 1 October 2020 to 10 March 2021
Hospitalisations
PHE undertook a record linkage to analyse sequenced confirmed VOC 202012/01 (B.1.1.7)
cases between October and December 2020. A Cox proportional hazard analysis of 63,609
sequenced cases identified a significantly increased risk of hospitalisation within 14 days of
specimen date among VOC 202012/01 (B.1.1.7) cases compared to non-variant cases (HR
1.34, 95% CI:1.07-1.66, p=0.01), after adjusting for confounders (sex, age, ethnicity, residence
type and week of diagnosis). This analysis used NHS hospital admission data from the NHS
Digital Secondary Uses Service (SUS) which is subject to reporting delays, therefore these
results represent a minimum estimate of hospital admission risk.
Deaths
As of 10 March 2021, there were 2,858 deaths (within 28 days) in the 109,093 cases with
confirmed or probable VOC 202012/01 (B.1.1.7) (case fatality 2.6%).
Immunisation
Two interim assessments have been published covering B.1.1.7 and the UK vaccine
programme.1
1 Early effectiveness of COVID-19 vaccination with BNT162b2 mRNA vaccine and ChAdOx1 adenovirus vector vaccine on symptomatic disease, hospitalisations and mortality in older adults in the UK: a test negative case control study Jamie Lopez Bernal et al.
2 Effectiveness of BNT162b2 mRNA Vaccine Against Infection and COVID-19 Vaccine Coverage in Healthcare Workers in England, Multicentre Prospective Cohort Study (the SIREN Study) Hall et al.
International Epidemiology
As of the 08 March 2021, there are 113 countries or territories (including the UK) reporting
cases of the UK variants globally. Of countries/territories outside of the UK, 19 report, or there is evidence of, community transmission. However for many countries the information available on the extent of transmission within the country is not clear.
GISAID (gisaid.org) includes data on sequences available internationally; as of the 10 March
2021 27,003 VOC 202012/01 (B.1.1.7) sequences (excluding UK) are available from 94
countries/territories.
VOC 202102/02 (B.1.1.7 cluster with E484K)
Through routine scanning of variation in VOC 202012/01 (B.1.1.7) a small number of B.1.1.7
sequences (65 of 135,310 sequences as of 10 March 2021), had acquired the spike protein
mutation E484K. Information suggested more than one independent acquisition event and on
investigation, this forms one predominant cluster and several separate cases or small clusters.
The predominant cluster was designated VUI on detection and on review re-designated as VOC
202102/02 (B.1.1.7 cluster with E484K) on 5 February 2021. The genomic and biological profile
is as previously described.
Epidemiological profile
As of 10 March 2021, 34 genomically confirmed cases of VOC 202012/01 (B.1.1.7) have been
identified; 21 with an epidemiological link to the South West and an additional 13 cases resident elsewhere in England (Table 4). Specimen dates range from 17 December 2020 to 20 February 2021, confirmed and probable cases by specimen date are shown in Figure 3.
| Region | Confirmed | Probable | Total | Percent of England total |
| London | 1 | 0 | 1 | 2.9 |
| North West | 11 | 0 | 11 | 32.4 |
| South East | 1 | 0 | 1 | 2.9 |
| South West | 21 | 0 | 21 | 61.8 |
| Total | 34 | 0 | 34 | 100.0 |
Figure 3. Confirmed and probable VOC 202102/02 (B.1.1.7 cluster with E484K) cases by specimen date, from 17 December 2020 as of 10 March 2021. Larger plot includes last 60 days only.
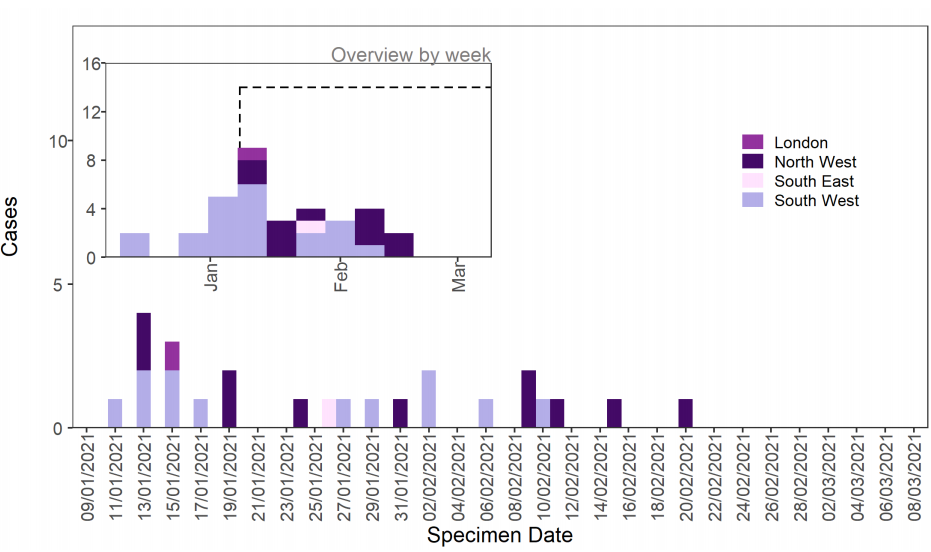
Deaths
As of 10 March 2021, one death (within 28 days) among 34 cases with confirmed VOC
202102/02 (B.1.1.7 cluster with E484K) has been reported.
International Epidemiology
As of the 8 March 2021, international cases have been reported in 2 countries and 2
sequences from the Netherlands have been identified on GISAID.
Variant in monitoring B.1.1.7 with S494P Clade
On 17 February 2021 the acquisition of Spike mutation S494P in VOC 202012/01 (B.1.1.7)
samples was identified as a signal as part of the horizon scanning process for routine
scanning of variation. Due to the large number of sequences identified, further analysis was
undertaken. This mutation appears to have been acquired multiple times by VOC 202012/01
(B.1.1.7). However, the majority of these sequences position within one clade, for which
further information is presented here.
Genomic and biological profile
The cluster VOC 202102/02 (B.1.1.7) cluster with S494P has the mutations previously
described for VOC 202012/01 (B.1.1.7) in technical briefing 6 with the addition of S494P in
spike gene and 2 mutations in orf1ab (NSP8 Q24Rand NSP5 SNP G10870T). S494P allows
evasion of binding or neutralization by several monoclonal antibodies but has not been
shown to impact on neutralization by convalescent or vaccine-induced polyclonal antisera.
S494P confers increased binding to hACE2.2
23Broad and potent activity against SARS-like viruses by an engineered human monoclonal antibody Rappazzo et al., 2021 Science DOI: 10.1126/science.abf4830
4Whelan Landscape analysis of escape variants identifies SARS-CoV-2 spike mutations that attenuatemonoclonal and serum antibody neutralization Liu et al., 2020. bioRxiv 2020.11.06.372037
Epidemiological profile
As of 10 March 2021, there were 761 genomically confirmed cases. Cases of VOC
202012/01 (B.1.1.7) with S494P have been increasing in recent weeks.
The distribution of cases over time shows that the clade began in the South East in
November 2020 and spread throughout this area in December 2020. It was also seen in the
North West during December 2020, expanding in January 2021.
VOC 202012/02 (B.1.351)
B.1.351 was initially detected in South Africa. This variant was designated VUI on detection
and on review re-designated as VOC 202012/02 (B.1.351) on 24 December 2020.
Epidemiological profile
VOC 202012/02 (B1.351) is dispersed across the UK in low numbers. Confirmed cases are
those identified by whole genome sequencing; probable cases are COVID-19 cases without
sequencing, but who are contacts of confirmed cases. As of 10 March 2021, 266 cases of
VOC 202012/02 (B.1.351) were identified (190 confirmed cases and 76 probable cases). An
international travel link was identified for 181 cases, and 68 cases had no travel link (17
awaiting further information). Confirmed and probable cases by specimen date are shown in
Figure 4, age-sex pyramid in Figure 5, and regional breakdown in Table 5.
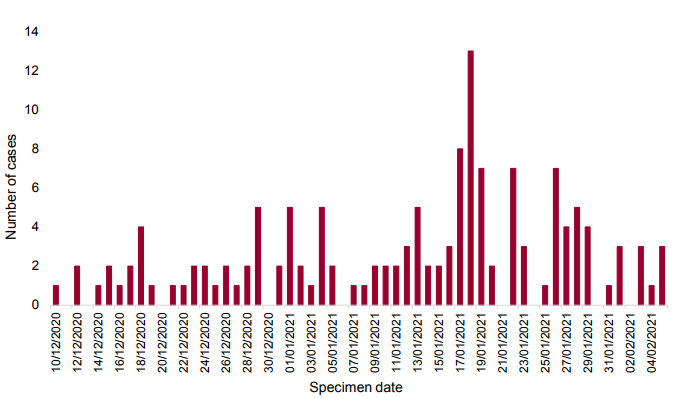
Figure 5. Age-sex pyramid of VOC 202012/02 (B.1.351) cases, from 10 December 2020 as of 10 March 2021
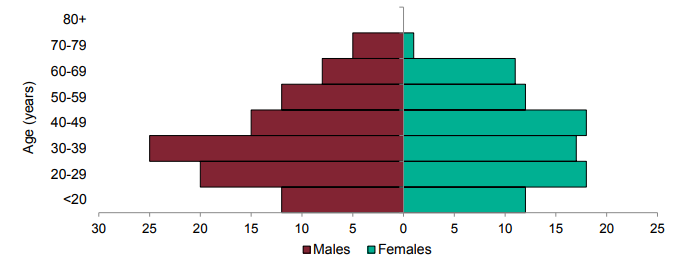
| Region | VOC 202012/02 (B.1.351) | |
| n | % | |
| East Midlands | 6 | 2.3 |
| East of England | 40 | 15.0 |
| London | 83 | 31.2 |
| North East | 6 | 2.3 |
| North West | 32 | 12.0 |
| South East | 54 | 20.3 |
| South West | 13 | 4.9 |
| West Midlands | 22 | 8.3 |
| Yorkshire and Humber | 10 | 3.8 |
| TBC | 0 | 0.0 |
| Total | 266 | |
Deaths
As of 10 March 2021, there were 6 deaths (within 28 days) in the 266 cases with confirmed or
probable VOC 202012/02 (B.1.351) (case fatality 2.3%).
International Epidemiology
As of 8 March 2021 there are 62 countries (including the UK) that have reported cases of this
variant globally. As of 08 March 2021 the epidemiological profile in South Africa is as follows:
• The case incidence is continuing to decrease. Currently, the reported weekly
incidence is 12.9 per 100,000 population. Weekly test positivity has also been
decreasing with current test positivity of 4.3%. Testing rates have been declining
since mid-January although remained relatively stable over the previous 2 weeks
(current weekly testing rate is 3.0 per 1,000 population).
• The fatality rate is decreasing (the weekly fatality rate is 1.2 per 100,000
population).
• The number of patients in hospital and ICU has also reduced slightly. As of 08
March 2021, 914 COVID-19 patients are in ICU and 6,105 are in hospital.
GISAID (gisaid.org) includes data on sequences available internationally. As of the 10 March
2021 2,643 sequences of VOC 202012/02 (B.1.351), excluding UK, are available from 46
countries/territories.
VOC 202101/02 (P.1)
First identified in Japan amongst travellers from Brazil, the P.1 lineage is a descendant of
B.1.1.28. This variant was designated VUI on detection and on review re-designated as VOC
202101/02 (P.1) on 13 January 2021. The clinical risk assessment for P1 is shown in Table 6.
| Indicator | RAG | Confidence | Assessment and rationale |
| Zoonotic emergence | NA | Not applicable | |
| Transmissibility between humans | MOD | Transmissibility appears greater than wild type virus in the context of Manaus, and there is MODERATE confidence that this represents a true change in phenotype. | |
| Infection severity | NA | Insufficient information. Reports from Manaus are confounded by healthcare system stresses. Clinical cohort studies currently underway in Brazil may provide data in the medium term. | |
| Naturally acquired immunity | HIGH | There is experimental evidence of evasion of naturally acquired immunity (HIGH confidence). There was a severe second wave in Manaus, despite apparently high seroprevalence. Better quality evidence, for example comparing the rate of confirmed reinfections in P1 and non P1 cohorts is required to raise the risk to red. | |
| Vaccine derived immunity | MOD | There is experimental evidence of evasion of vaccine derived immunity in laboratory studies (MODERATE confidence). The magnitude of the effect may be less than that of B.1.351 (South Africa), based on limited laboratory data . Clinical trial and surveillance data are required. | |
| Drugs and Therapeutics | MOD | There is experimental evidence that variants containing E484K may have reduced susceptibility to some monoclonal antibody based therapies in laboratory conditions (MODERATE confidence). | |
| Overall assessment of level and nature of risk, and level of confidence | There are limited data available on P.1. It is a successful lineage in Brazil and is showing some evidence of international spread. The spread of P1 in the UK, where the prevalent virus is B.1.1.7, cannot be extrapolated from the trajectory observed in Manaus. There is laboratory evidence supporting antigenic change, and this is consistent with the epidemiology reported from Brazil. In the context of the situation in Manaus, it is difficult to differentiate immune escape and increased transmissibility, and some degree of both is plausible. . However, there are no robust, comparative clinical re-infection or vaccine efficacy data. There is an urgent need to assess severity. |
Epidemiological profile
As of 10 March 2021, 7 genomically confirmed and probable cases of VOC 202101/02 (P.1) have been identified in England. All cases have been linked to international travel.
Deaths
As of 10 March 2021, there were no reported deaths in cases of VOC 202101/02 (P.1) in England.
International Epidemiology
As of 8 March 2021, cases of VOC 202101/02 (P.1) have been reported in 20 countries or territories. 8 countries have reported cases of a Brazilian variant additional information is awaited to clarify if this is with VOC 202101/02 (P.1).
GISAID (gisaid.org) includes data on sequences available internationally. As of the 10 March 2021 562 sequences of VOC 202101/02 (P.1) are listed (Brazil 304, Italy 102, Belgium 29, Peru 24, USA 18, Colombia 17, Switzerland 16, Portugal 11, France 9, Japan 8, Ireland 6, Netherlands 4, French Guiana 3, Romania 2, South Korea 1, Canada 1, Mexico 1, Spain 1, Sint Maarten 1, Germany 1, Turkey 1, Australia 1, Faroe Islands 1)
VUI 202101/01 (P2)
First identified in Brazil, the P.2 lineage is a descendant of B.1.1.28. This variant was designated VUIon detection and on review re-designated as VUI 202101/01 (P.2) on 13 January 2021.
Genomic profile
VUI 202101/01 is lineage P.2 (first sequence detected in Brazil and was first sequenced in the UK in November 2020). The complete mutation profile of VUI 202101/01 (P.2) is shown in Table 7 and genomic case definition in Table 8.
| Gene | Amino Acid | Actual Nucleotide | Note |
| - | 100>T | not included in definition due to masking | |
| orf1ab | L3468V | 10667T>G | nsp5:L205V |
| - | 11824C>T | nsp6:I248I | |
| - | 12964A>G | nsp9:G93G;not lineage defining | |
| S Gene | E484K | 23012G>A | |
| orf8 | F120F | 28253C>T | |
| N Gene | A119S | 28628G>T | |
| M234I | 28975G>T | ||
| - | 29754>T | not included in definition due to masking |
| CONFIRMED | All variant defining changes called as alternate base or 6 of 7 changescalled as alternate base and remaining position either N or mixed base |
| PROBABLE | NA |
| LOW_QC | Fewer than 6 positions are called but at least one is called as alternate (variant) base and all other defining positions reported as N (unknown) or mixed bases. |
Epidemiological profile
As of 10 March 2021, 46 cases of VUI 202101/01 (P.2) have been identified in England. 9 cases
have been linked to international travel, and 32 cases had no travel link (5 awaiting further
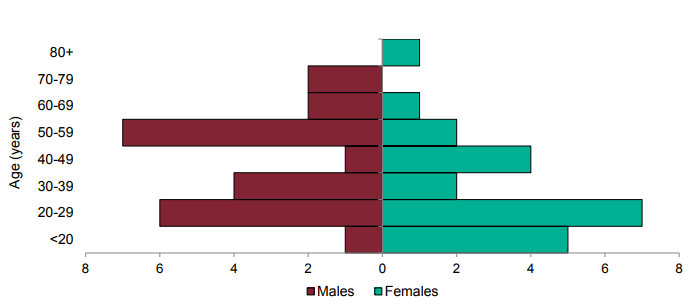
information). Confirmed and probable cases by specimen date are shown in Figure 6 and agesex
pyramid in Figure 7.
Figure 7. Age-sex pyramid of VUI 202101/01 (P.2) cases, 10 December 2020 to 10 March 2021
International Epidemiology
As of 8 March 2021, cases of VUI 202101/01 (P.2) have been reported in 11 countries or territories.
GISAID (gisaid.org)includes data on sequences available internationally. As of the 10 March 2021 862 sequences (excluding UK) of VUI 202101/01 (P.2) are listed ( Brazil 382, USA 267, French Guiana 42, Canada 23, Switzerland 22, Netherlands 21, Denmark 18, Portugal 16, Spain 11, Ireland 11, France 10, Norway 6, Chile 6, Argentina 5, Germany 4, Australia 3, Japan 3, Sweden 3, Mexico3, Singapore 1, Austria 1, Italy 1, Sint Maarten 1, Belgium 1, Malaysia 1).
VUI 202102/01 (A.23.1 with E484K)
This variant was first identified in Liverpool, UK, derived from a lineage first identified in Uganda without E484K. The variant was designated VUI on detection and on review re-designated as VUI 202102/01 (A.23.1 with E484K) 5 February 2021.
Genomic profile
VUI 202102/01 (A.23.1 with E484K) is lineage A.23.1 (first sequence detected in the UK in
December 2020). The complete mutation profile of VUI 202102/01 (A.23.1 with E484K) is shown
in Table 9 and genomic case definition in Table 10.
| Gene | Amino Acid | Actual Nucleotide | Note |
| orf1ab | L1559F | 4940C>T | nsp3:L741F |
| M3655I | 11230G>T | nsp6:M86I | |
| L3667F | 11266G>T | nsp6:L98F | |
| M3752I | 11521G>T | nsp6:M183I | |
| S Gene | R102I | 21867G>T | |
| F157L | 22033C>A | ||
| V367F | 22661G>T | ||
| E484K | 23012G>A | ||
| Q613H | 23401G>T | ||
| P681R | 23604C>G | ||
| orf8 | L84S | 28144T>C | |
| N Gene | E92K | 28167G>A | |
| S202N | 28878G>A |
| CONFIRMED | All variant defining changes called as alternate base |
| PROBABLE | At least 5 variant defining changes called as alternate base and all other positions either N or mixed base |
| LOW_QC | Fewer than 5 variant defining changes called as alternate base and all other positions either N or mixed base |
Epidemiological profile
As of 10 March 2021, 59 genomically confirmed and 19 genomically probable cases of VUI
202102/01 (A.23.1 with E484K) have been identified; 78 in total. The majority of these are
residents of the North West of England (Table 11). Specimen dates range from 26 December
2020 to 23 February 2021 (Figure 8).
| Region | Confirmed | Probable | Total | Percent of England total |
| North West | 58 | 19 | 77 | 98.7 |
| West Midlands | 1 | 0 | 1 | 1.3 |
| Total | 59 | 19 | 78 | 100.0 |
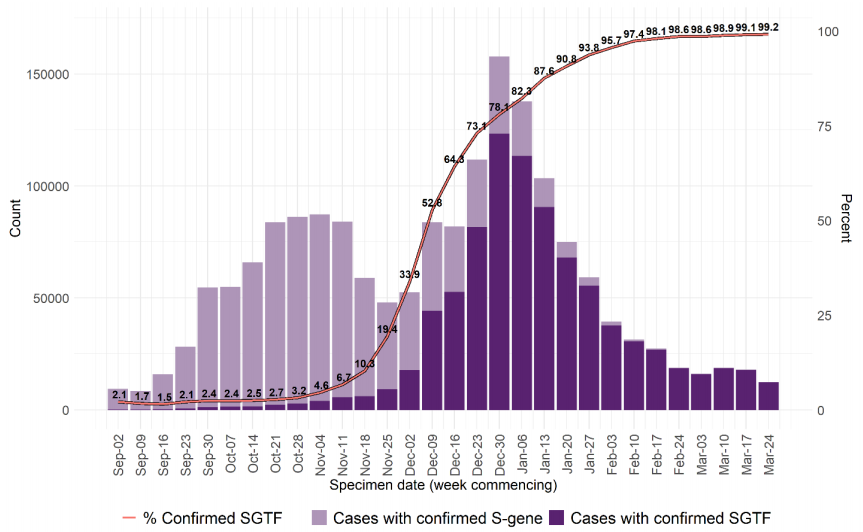
Deaths
As of 10 March 2021, 1 death (within 28 days) in 78 cases with VUI 202102/01 (A.23.1 with
E484K) has been reported (case fatality 1.3%).
International Epidemiology
The are no cases reported internationally as of the 8 March 2021.
GISAID (gisaid.org) includes data on sequences available internationally. As of the 10 March
2021 1 sequence is listed of VUI 202102/01 (A.23.1 with E484K) (excluding UK) from
Netherlands.
VUI 202102/03 (B.1.525)
First identified as a geographically dispersed cluster in UK on the 2 February 2021. This variant was designated VUI on detection and on review re-designated as VUI 202102/03 (B.1.525) on 12 February 2021.
Genomic profile
VUI 202102/03 (B.1.525) is lineage B.1.525 (first sequence detected in the UK in February
2021). The complete mutation profile of VUI 202102/01 (A.23.1 with E484K) is shown in Table
12 and genomic case definitions in Table 13.
| Gene | Amino Acid | Actual Nucleotide | Note |
| orf1ab | - | 1498C>T | nsp2:F231F |
| - | 1807A>G | nsp2:G334G | |
| - | 2659G>A | ||
| T2007I | 6285C>T | nsp3:T1189I | |
| - | 8593T>C | nsp4:V13V | |
| - | 9565C>T | nsp4:F337F | |
| 3675_7del | 11288_96del | nsp6:106_8del | |
| P4715S | 14407C>T | nsp12:P323S | |
| - | 18171C>T | nsp14:G44G | |
| - | 20724A>G | nsp16:L22L | |
| S Gene | Q52R | 21717A>G | |
| A67V | 21762C>T | ||
| 69_70del | 21765_70del | ||
| 144del | 21991_3del | ||
| E484K | 23012G>A | ||
| Q677H | 23593G>C | ||
| F888L | 24224T>C | ||
| - | 24748C>T | F1062F | |
| E Gene | L21F | 26305C>T | |
| M Gene | I82T | 26767T>C | |
| orf6 | 2del | 27205_7del | |
| N Gene | 2_3del | 28278_80del | |
| A12G | 28308C>G | ||
| - | 28699A>G | P142P | |
| - | 29543G>T |
Red text indicates acquisition in subset of isolates within the lineage, non-variant defining
mutations. Indels (shaded in orange) are not currently included in variant definitions
| CONFIRMED | All variant defining changes called as alternate base |
| PROBABLE | At least 5 variant defining changes called as alternate base and all other positions either N or mixed base |
| LOW_QC | Fewer than 5 variant defining changes called as alternate base and all other positions either N or mixed base |
Biological profile
The significance of E484K has been described previously in this briefing. Q677H is proximal to
the furin cleavage site, it is unclear what the impact of this mutation might be, however
mutations around the furin cleavage site are known to modulate transmissibility. The deletions
in the NTD (Δ69-70 and Δ144) are also found in the VOC 202012/01
(B.1.1.7) variant, Δ144 is an antigenic escape mutant and is highly associated with virus
replication in immunocompromised patients. The deletion in NSP6 (Δ106-108) is found in
multiple variants of concern (VOC 202012/01 (B.1.1.7) variant, VOC 202012/02 (B.1.351) and
VOC 202101/02 (P.1) variants). NSP6 is involved in immune evasion but the impact of this
deletion is currently unclear.
Deaths
As of 10 March 2021, 4 deaths in 107 cases with VUI 202102/03 (B.1.525) has been reported
(case fatality 3.7%).
International Epidemiology
As of 8 March 2021, cases of VUI 202102/03 (B.1.525) have been reported in 11 countries or
territories
GISAID (gisaid.org) includes data on sequences available internationally. As of the 10 March
2021 328 sequences of VUI 202102/03 (B.1.525) are listed, excluding UK. (Denmark 120,
Nigeria 76, USA 42, Italy 15, Slovenia 41, Canada 8, Japan 7, Netherlands 7, Germany 7,
Ireland 6, France 6, Norway 4, Belgium 3, Australia 2, Jordan 2, Cameroon 2, Malaysia 2,
Switzerland 2, Austria 2, Singapore 1, Finland 1, Thailand 1, Mayotte 1, Spain 1).
VUI 202102/04 (B.1.1.318)
The VUI 202102/04 (B.1.1.318) was identified in England in mid February 2021 through routine
horizon scanning for the development of new clusters within the E484K genomes. This analysis
identified an initial cluster of 6 cases containing E484K and other spike mutations, designated
VUI 202102/04 (B.1.1.318) 23 February 2021.
Genomic profile
VUI 202102/04 (B.1.1.318) is lineage B.1.1.318. The complete mutation profile of VUI
202102/04 (B.1.1.318) is shown in Table 14 and genomic case definitions in Table 15.
| Gene | Amino Acid | Actual Nucleotide | Note |
| orf1ab | E378V | 3852A>T | nsp3:E378V |
| - | 3961C>T | ||
| K2511N | 7798G>T | nsp3K1693N | |
| T2936I | 9072C>T | nsp4:T173I | |
| A3209V | 9891C>T | nsp4:A446V | |
| T3284I | 10116C>T | nsp5:T21I | |
| 3675_7del | 11288_96del | nsp6:106_8del | |
| V6672M | 20578G>A | nsp15:V320M | |
| S | T95I | 21846C>T | |
| 144del | 21991_3del | ||
| E484K | 23012G>A | ||
| - | 23287T>C | ||
| P681H | 23604C>A | ||
| D796H | 23948G>C | ||
| - | 24382C>T | ||
| - | 25276C>A | ||
| M | I82T | 26767T>C | |
| orf8 | 1_3del | 27894_901del | |
| E106 | 28209G>T | ||
| - | 28271A>G | ||
| N | A208_A209delinsG | 28896_8del |
| CONFIRMED | All variant defining changes called as alternate base |
| PROBABLE | At least 5 variant defining changes called as alternate base and all other positions either N or mixed base |
| LOW_QC | Fewer than 5 variant defining changes called as alternate base and all other positions either N or mixed base |
Epidemiological profile
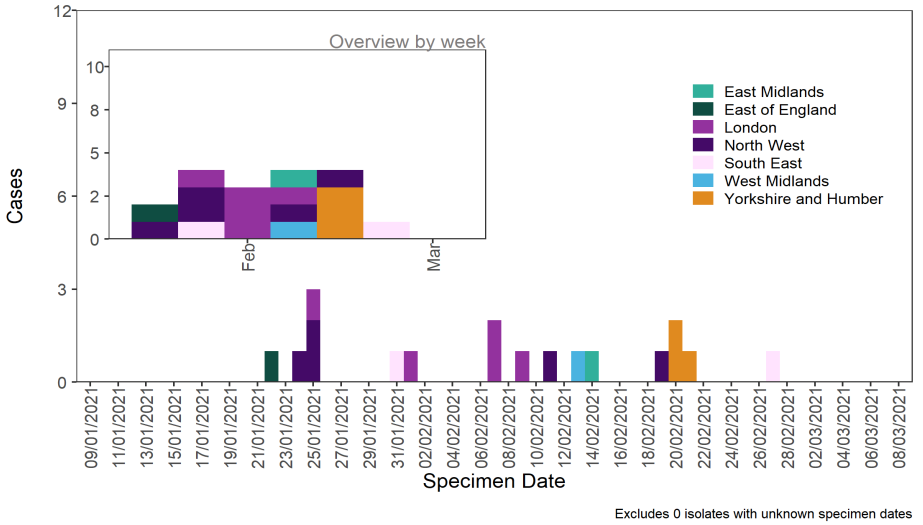
As of 10 March 2021, there were 18 genomically confirmed cases of VUI 202102/04 (B.1.1.318).
Cases are based across England (Table 16), with specimen dates ranging from 22 January 2021
to 27 February 2021 (Figure 9). Three of the initial 6-person cluster are related to cases that
travelled to Nigeria, however the wide geographic spread suggests this VUI may be a result of
multiple importations or could be an undersampled within the UK.
| Region | Confirmed | Probable | Total | Percent of England total |
| East Midlands | 1 | 0 | 1 | 5.6 |
| East of England | 1 | 0 | 1 | 5.6 |
| London | 5 | 0 | 5 | 27.8 |
| North West | 5 | 0 | 5 | 27.8 |
| South East | 2 | 0 | 2 | 11.1 |
| West Midlands | 1 | 0 | 1 | 5.6 |
| Yorkshire and Humber | 3 | 0 | 3 | 16.7 |
| Total | 18 | 0 | 18 | 100.2 |
Deaths
No deaths (within 28 days of specimen date) among confirmed or probable VUI 202102/04
(B.1.1.318) have been reported as of 10 March 2021.
International Epidemiology
As of 8 March 2021 there are no cases reported internationally.
GISAID (gisaid.org) includes data on sequences available internationally. As of the 10 March
2021, there are no international VUI 202102/04 (B.1.1.318) sequences
VUI 202103/01 (B.1.324.1 with E484K)
First identified via horizon scanning of genomes with spike mutations characteristic of VOCs
(including both N501Y and E484K) on 3 March 2021, the variant VUI 202103/01 (B.1.324.1 with
E484K) was designated VUI on detection as VUI 202103/01 (B.1.324.1 with E484K) on 4 March
2021.
Genomic profile
The complete mutation profile of VUI 202103/01 (B.1.324.1 with E484K) is shown in Table 17
and genomic case definitions in Table 18.
| Gene | Amino Acid | Nucleotide | Note |
| orf1ab | T265I | 1059C>T | nsp2:T85I |
| G894S | 2945G>A | nsp3:G76S | |
| - | 3037C>T | ||
| - | 6730C>T | ||
| T2196P | 6851A>C | nsp3:T1378P | |
| - | 8704T>C | from wider lineage | |
| - | 8986C>T | ||
| T2952I | 9120C>T | nsp4:T189I | |
| T3202M | 9870C>T | nsp4:T439M | |
| H3580Q | 11005C>A | nsp6:H11Q | |
| P4197S | 12854C>T | nsp9:P57S | |
| S4398L | 13458C>T | nsp12:S6L | |
| - | 13617G>A | from wider lineage | |
| P4715L | 14408C>T | nsp12:P323L | |
| - | 14559G>A | ||
| G5530C | 16852G>T | nsp13:G206C | |
| S Gene | - | 22388C>T | |
| E484K | 23012G>A | ||
| S494P | 23042T>C | ||
| N501Y | 23063A>T | ||
| D614G | 23403A>G | ||
| P681H | 23604C>A | ||
| E1111K | 24893G>A | ||
| orf3a | Q57H | 25563G>T | |
| orf8 | 10_21del | 27922_56del | |
| - | 28272A>T | from wider lineage | |
| NGene | M234I | 28975G>A |
| CONFIRMED | All 8 variant defining changes called as alternate base |
| PROBABLE | NA |
| LOW_QC | Fewer than 8 variant defining changes called as alternate base and all other positions either N or mixed base |
Epidemiological profile
As of 10 March 2021, there are 2 confirmed cases in the UK in a single group of returning
travellers. Two additional households note contact of which one has tested positive, sequence
data not available. There is no current evidence of spread in the UK based on sequencing data.
No deaths were reported.
International Epidemiology
As of 8 March 2021 there are no cases reported internationally.
GISAID (gisaid.org) includes data on sequences available internationally. As of the 10 March
2021, 0 sequences are listed internationally of VUI 202103/01 (B.1.324.1 with E484K).
Appendices
Appendix 1. SGTF Surveillance
One S gene mutation in VOC 202012/01 (B.1.1.7) causes deletion of amino acids 69 and 70
(Δ69-70), with a reproducible S gene target failure (SGTF). This is detected by the
ThermoFisher TaqPath assay used in UK lighthouse laboratories (see Technical Briefing 1),
‘TaqPath laboratories.’
Considering pillar 2 samples where we know both the sequence and the SGTF status, 99.6% of
Δ69-70 sequences are SGTF, compared to 0.05% of sequences without the deletion. Since 1
January 2021, > 99.7% of Δ69-70 sequences are VOC 202012/01 (B.1.1.7) in all regions of
England.
Surveillance of SGTF, as a proxy for VOC 202012/01 (B.1.1.7), is based on positive tests
reported by 3 lighthouse laboratories that use the Thermo Fisher TaqPath RT-PCR, and for
which CT values are low enough to classify if the S gene is detectable. Specifically, positive
tests with CT values >30 for any gene target are excluded. SGTF is defined as a positive test
with CT values <=30 for the N and ORF1ab genes and an undetectable S gene. S gene positive
is a positive test for which all 3 gene targets (N, ORF1ab, S) have CT values <=30.
Samples with SGTF have predominated since mid-December 2020, and has remained above
95% since the beginning of February 2021 (Figure 10). All regions in England are >98% SGTF
in the most recent week (3 March to 9 March 2021; Figure 11). Total cases detected using the
TaqPath assay have also been declining since the first week of 2021, reflecting the general
decline in case rates across England.
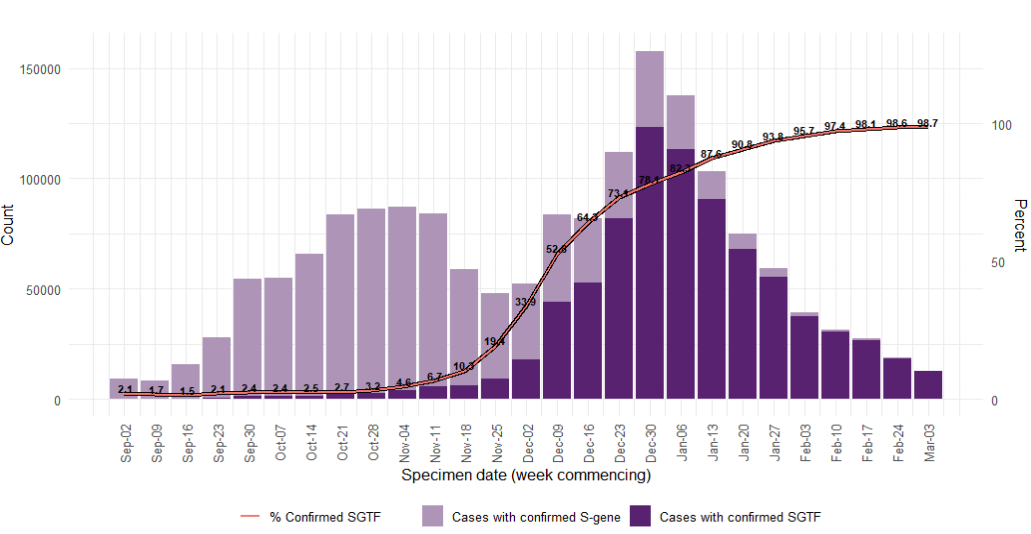
Figure 11. Weekly number and proportion of England Pillar 2 COVID-19 cases with SGTF among those tested with the TaqPath assay and with S gene detection results, by region of residence (1 September 2020 to 9 March 2021)
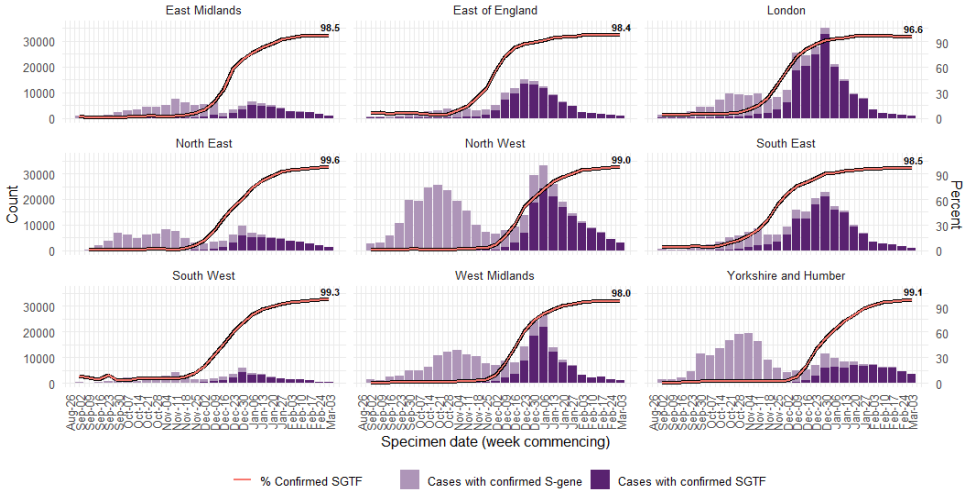
Data on coverage of TaqPath laboratories testing and numbers/proportions of cases with SGTF are shared daily with Local Authorities (Sunday to Friday) on the COVID-19 PHE Local Authorities Report Store (Sharepoint).
Sources and acknowledgments
Data sources
Data used in this investigation is derived from the COG-UK dataset, the PHE Second
Generation Surveillance System (SGSS), NHS Test and Trace, the Secondary Uses Service
(SUS) dataset and Emergency Care Data Set (ECDS).
Variant Technical Group
Organisations
This group includes representation from the following organisations: PHE, DHSC, BEIS, Wales
NHS, PHScotland, NHS Scotland, Health and Social Care Northern Ireland, Imperial College
London, London School of Hygiene and Tropical Medicine, University of Birmingham, University
of Cambridge, University of Edinburgh, University of Liverpool, the Wellcome Sanger Institute.
Acknowledgements
The authors are grateful to those teams and groups provid
ing data for this analysis, including: the Lighthouse Laboratories, COG-UK, the Wellcome
Sanger Institute, the PHE Epidemiology Cell, Contact Tracing, Genomics and Outbreak
Surveillance Teams.
Published February 2021
PHE gateway number: GW-1934






