Spike whole mutant SARS-CoV-2(2019nCoV) 501Y.V2 lineage(B.1.351) Pseudotyped virus production and Pseudovirus Based Neutralization Assay
Cat No.:GM-2019nCoV-PSV20
Order information
| Catalot Number | Size (T,Test, formalized in 96 well, one test per well) |
Price(In USD) | Qty (Quantity) | Sum(In USD) |
|---|---|---|---|---|
| GM-2019nCoV-PSV20 | 100T | 3840 | ||
| GM-2019nCoV-PSV20 | 200T | 5440 | ||
| GM-2019nCoV-PSV20 | 1000T | 21760 | ||
| GM-2019nCoV-PSV20 | 5000T | 87040 | ||
| For larger scale | Click for inquiry. | Click for inquiry. | ||
| Shipping Cost: | 960.00 | |||
| Total: | ||||
GeneMedi pseudotyped virus (pseudovirus) of SARS-COV-2 (2019nCOV) 501Y.V2 lineage
|
Background Reading: |
The world is in midst of the COVID-19 pandemic. Recently a new SARS-CoV-2 (2019nCOV)lineage (501Y.V2,B.1.351), characterised by eight lineage-defining mutations in the spike protein (excluding D614G mutation), including three at important residues in the receptor-binding domain (K417N, E484K and N501Y), represents increased transmissibility. This lineage emerged in South Africa and spread rapidly, becoming within weeks the dominant lineage in the Eastern Cape and Western Cape Provinces of South Africa.
Mutations of Spike protein in SARS-COV-2 (2019nCOV) 501Y.V2 lineage(B.1.351) (B.1.351) spread in South Africa
Three of the spike mutations are at key residues in the RBD (N501Y, E484K and K417N), three are in the N-terminal domain (L18F, D80A and D215G) and one is in loop 2 (A701V). Deletion of three amino acids at L242-244L is disputed with a L242H mutation1.
| Spike Mutation in SARS-COV-2 (2019nCOV) 501Y.V2 lineage(B.1.351) |
Spike-S1 Subunit N Terminal |
L18F | |
| D8OA | |||
| D215G | |||
| Spike-S1 Subunit | L242-244 deletion | disputed with a L242H mutation | |
| Spike-S1 Subunit | R246I | ||
| Spike-RBD | K417N | Important residues mutation | |
| E484K | |||
| N501Y | |||
| Spike-S1 Subunit | D614G | A popular mutation in different new SARS-CoV-2 lineage | |
| Spike-S2 Subunit | A701V |
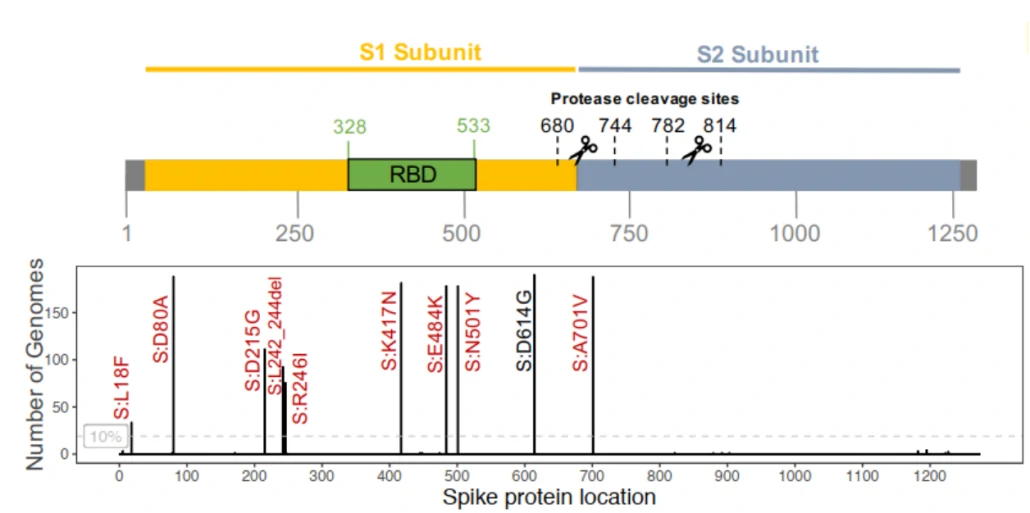
Figure. Amino acid changes in the spike region of the 190 S501Y.V2 genomes1








 PSV validated Data Poster Download
PSV validated Data Poster Download