Adenovirus-LC3 production service for Autophagy Flux Detection
Order Information
| Product | Adv Vector Production Scale | Price(In USD) | Qty (Quantity) | Sum(In USD) |
|---|---|---|---|---|
| Ad-mRFP-GFP-LC3 | 1E+10PFU | 1880 | ||
| Ad-mRFP-GFP-LC3 | 1E+11PFU | 5400 | ||
| Ad-mRFP-GFP-LC3 | 1E+12PFU | 23000 | ||
| Ad-GFP-LC3 | 1E+10PFU | 1880 | ||
| Ad-GFP-LC3 | 1E+11PFU | 5400 | ||
| Ad-GFP-LC3 | 1E+12PFU | 23000 | ||
| Shipping Cost: | 760.00 | |||
| Total: | ||||
Introduction to Adenovirus-LC3 production service for Autophagy Flux Detection
Premade LC3 Autophagy Biosensors Products and user manual
Adeno associated virus(AAV) AAV-GFP-LC3 Autophagy BiosensorAAV-mRFP-GFP-LC3 Autophagy Biosensor |
Adenovirus Adv-GFP-LC3 Autophagy BiosensorAdv-mRFP-GFP-LC3 Autophagy Biosensor |
Lentivirus Lv-GFP-LC3 Autophagy BiosensorLv-mRFP-GFP-LC3 Autophagy Biosensor |
LC3 Autophagy Biosensors User Manual
AAV-LC3 User Manual  |
Adenovirus-LC3 User Manual  |
Lentivirus-LC3 User Manual  |
Autophagy is a highly regulated homeostatic degradative process where cells destroy their own components via the lysosomal machinery and recycle them. This process plays both protective and deleterious roles in many diseases, including Alzheimer’s disease, aging, cancer, infection and Crohn’s disease. Elucidating the correlation between autophagy and apoptotic cell death has become the focus of a great deal of research. Members of the LC3 family play a key role in the maturation of the autophagosome. Lysosomal turnover of the autophagosomal marker LC3-II reflects starvation-induced autophagic activity, and detecting LC3 by immunoblotting or immunofluorescence has become a reliable method for monitoring autophagy and autophagy-related processes, including autophagic cell death.
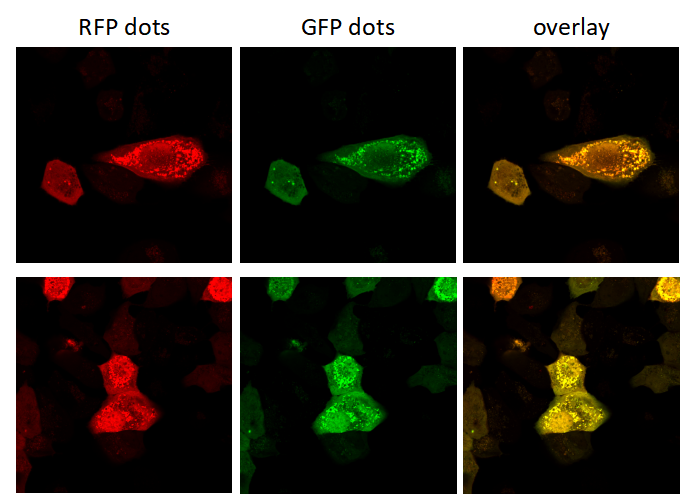
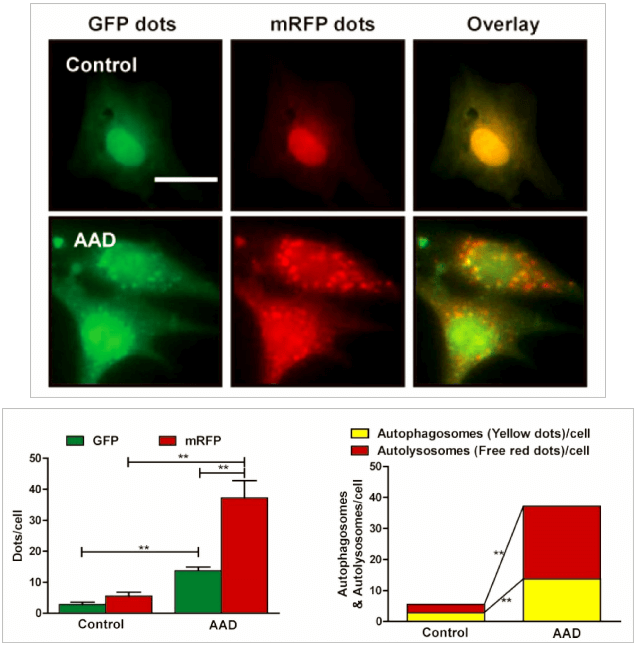
Genemedi has launched series of Adenovirus packaging service of autophage related biosensors, in which GFP and/or RFP tags are fused at the C-termini of the autophagosome marker LC3, allowing to detect the intensity of autophagy flux in real-time with more accuracy, clarity and intuitiveness. These biosensors provides an enhanced dissection of the maturation of the autophagosome to the autolysosome, which capitalizes on the pH difference between the acidic autolysosome and the neutral autophagosome. The acid-sensitive GFP will be degraded in autolysosome whereas the acid-insensitive RFP will not. Therefore, the change from autophagosome to autolysosome can be visualized by imaging the specific loss of the GFP fluorescence, leaving only red fluorescence. Taking advantage of RFP-GFP-LC3 and GFP-LC3 labeling system, Genemedi has launched the production service of Adv-RFP-GFP-LC3 and Adv-GFP-LC3, which can be used to observe autophagy flux and monitor the intensity of autophagy flux in real-time in vivo or in vitro.
Properties
| Adenovirus Vector | |
|---|---|
| Quantity/Unit | Vials |
| Form | Frozen form |
| Sipping and Storage Guidelines | Shipped by dry ice, stored at -80 ° C, effective for 1 year. Avoid repeatedly freezing and thawing |
| Titer | > 1*10^10 PFU/ml |
Advantages
1. Autophagy flux detection in real time, quantitative.
2. High resolution, more accuracy and sensitivity than traditional approaches.
3. Broad range of host. Adv has the ability to infect dividing, quiescent cells, stem cells, and primary cells, allowing genetic materials to be delivered to a highly diverse range of cell types and tissues.
4. High infection efficiency. Almost 100% gene delivery in most cell types, completely surpassing other viral vector tools and liposome transfection.
5. Great packaging capacity, up to 8kb.
6. Without integration. No alteration to host genome.
7. High titer. Up to 1011 pfu/ml.
Applications and Figures
Quality control description
Our optimized custom adenoviral vector production and strict quality control systems provide customers a high titer of functional recombinant adenoviral vectors. Viral titers are determined by TCID50 method, which is the most accurate way to measure the titer of adenovirus.
Technical Documents
1. For detailed protocol about how to package and purify Adv, please see the  Adenovirus User Manual on the Genemedi website.
Adenovirus User Manual on the Genemedi website.
2. For further information about Adv autophagy flux detection system, please refer to the  Adenovirus Autophagy Flux Detection Manual on the Genemedi website.
Adenovirus Autophagy Flux Detection Manual on the Genemedi website.
Frequently Asked Questions(FAQs)
- 1. Why does adenovirus have a relatively higher immunogenecity compared with rAAV?
-
Answer
The adenovirus without E1/E3 can express all of the other genes in the viral backbone and hence induces immunogenic responses, while rAAV does not have any of the AAV genes, thus no immunogenicity from viral protein.
- 2. What is the role of the E1 Gene in adenoviruses?
-
Answer
Simply, the E1 gene products are early proteins that are transcribed in the early transcribed regions and required for proceeding subsequent steps in viral replication. The E1 gene contains E1A and E1B, involved in the replication of adenovirus. E1A is critical to start viral replication by promoting transcription from rep gene promoters, P5 and P19, and facilitate viral replication by activating the early adenovirus promoters.
- 3. How to store adenovirus?
-
Answer
It would be better to store adenovirus in PBS at -80oC. Sucrose or DMSO may help to stabilize the vector.
- 4. How can you tell if your vector is lentiviral, retroviral, or adenoviral?
-
Answer
You should blast your vector sequence and see if there’re sequences of reverse transcriptase and integrase (gene names: gag and pol), which are for lentiviral/retroviral vectors, but not for adenoviral. For another, if your plasmid is around 30-35kb in size, it's certainly adenoviral.
- 5. Is adenovirus a useful tool to study primary macrophage functions?
-
Answer
In RAW264.7 and PM cells, adenovirus works very well, and it seems that IL1β expression is increased slightly after adenovirus transfection compared with negative control. While for the BMDM, adenovirus does not work, it may be better to use lentivirus instead, which gives a pretty good transduction efficacy and less inflammatory response.
- 6. Is it possible to infect a tissue preparation with lentivirus and afterwards with adenovirus and getting high efficiency in transduction?
-
Answer
It is definitively possible to perform sequential transductions/infections. Polybreen, protamine sulfate or other transduction enhancing reagents are recommended to enhance viral particle infectivity.
- 7. What is the difference between LC3A, LC3B and LC3C, or LC3-I and LC3-II?
-
Answer
LC3 is a soluble protein with a molecular mass of ∼17 kDa and is distributed ubiquitously in eukaryotes. It is expressed as the splice variants LC3A, LC3B, and LC3C which display unique tissue distribution. All LC3 isoforms undergo post-translational modifications, especially PE conjugation (lipidation) during autophagy. Upon autophagic signal, the cytosolic form of LC3 (LC3-I) is conjugated to PE to form LC3-PE conjugate (LC3-II), which is recruited to the autophagosomal membranes.
- 8. Can I use the Ad-mRFP-GFP-LC3 products to study autophagy in cells in combination with a lentivirus transfection?
-
Answer
Adenovirus transfection may actually slightly induce autophagy, skewing the results. Make certain to include appropriate controls and make sure to leave an appropriate time (48-72hrs) post transfection to lower the basal autophagy level.
- 9. Will the GFP signal of mRFP-GFP-LC3 that anchored at autophagosomes and fused with lysosome relight after the cell fixed?
-
Answer
GFP is no longer delighted reversibly once GFP-LC3 signal was destroyed by the acidic environment.
- 10. Can transient expression of RFP-GFP-LC3 differ badly from stable expression?
-
Answer
Transient transfection of GFP-LC3 can leads to the formation of non-autophagic LC3 puncta due to overexpression. Reports shows that GFP-LC3 is aggregate prone protein. We suggest the use of stable cell line for assessment of GFP-LC3 puncta.
- 11. Which software do you usually use for counting the number of LC3 dots in the cell?
-
Answer
Image J does work. For LC3 puncta, we suggest using the "Watershed" plugin from Image J. But proper threshold adjustment is critical. CellProfiler is another choice. First use the top hat filter. Then with the Identify Primary Objects module, manually adjust the Intensity threshold to optimally select the punctas of the right intensity.
- 12. How long is chloroquine half life when I treat a cell line for an autophagy study?
-
Answer
Chloroquine is an attractive drug agent effective for the treatment of not only malaria but also inhibition of autophagy, which is a promising effect for anti-tumor therapy. Half-life time of this drug is approximately 18 hours in vivo due to the degradation by the liver, but the stability of chloroquine in vitro experiments is expected to be longer. It will depend mainly on the cell line your are working on and their metabolic activity. For example HepG2 cells (human hepatocarcinoma cells) have a strong capacity to metabolize drugs. this might not be the case for other cell line of different tissular origin. I would suggest to add chloroquine when you change your growth medium.
- 13. What medium is best to use to induce LC3 puncta in HeLa cells?
-
Answer
We recommend the media used by Axe et al. (2008) JCB 182: 685-701. The recipe is 140 mM NaCl, 1 mM CaCl2, 1 mM MgCl2, 5 mM glucose, and 20 mM Hepes, pH 7.4, Add 1% BSA and pass through a 20um filter before use.
- 14. What is the best applicable inhibitor of autophagy?
-
Answer
I prefer to use CQ (chloroquine) in all of my researches about autophagy inhibition. The using of CQ is quite easy, since it is easy dissolve in water (unlike 3-MA in DMSO). One time I have been used 3-MA, actually I did not like to work with it. Bafilomycin is another choice for you.
- 15. When to add bafilomycin to study autophagy?
-
Answer
it is better to do a time course to be sure. We use at least two times (2 and 4 hours) to starved the cells and we add baf for 2 hours (for 4 hours: the 2 last hours). We then follow LC3-II and P62 by WB. To see the degradation of p62, sommetime 4 hours is better than 2hours.
- 16. Which neuronal cell line should I use for autophagy level?
-
Answer
SHSY5Y and PC12 are two well established neuronal cell lines, human and rat cell lines, respectively. You may also try Neuro2A, it is a mouse cell line.
- 17. How to distinguish selective and non-selective autophagy?
-
Answer
You can analyze selective mitophagy by interaction/co-localization study of mitochondrial marker protein and the autophagy specific adaptor protein LC3II.
- 18. When to add the autophagy treament in cell culture models?
-
Answer
First, some cells are very sensitive to these inhibitors and you need to optimize your conditions. The second point, if you like to stop autophagy process after inducing it to compare whether it is a cell death or cell survival you may add it just hours (2-3) before treatment. However, many articles prefer to use it as pre-treatment 1 hour before any other treatment.
Reference
1. Hariharan N, P Zhai and J Sadoshima. (2011). Oxidative stress stimulates autophagic flux during ischemia/reperfusion. Antioxid Redox Signal 14:2179-90.
2. Yuan Y, Y Zheng, X Zhang, Y Chen, X Wu, J Wu, Z Shen, L Jiang, L Wang, W Yang, J Luo, Z Qin, W Hu and Z Chen. (2017). BNIP3L/NIX-mediated mitophagy protects against ischemic brain injury independent of PARK2. Autophagy 13:1754-1766.
3. Yu T, F Guo, Y Yu, T Sun, D Ma, J Han, Y Qian, I Kryczek, D Sun, N Nagarsheth, Y Chen, H Chen, J Hong, W Zou and JY Fang. (2017). Fusobacterium nucleatum Promotes Chemoresistance to Colorectal Cancer by Modulating Autophagy. Cell 170:548-563 e16.
4. Shao BZ, P Ke, ZQ Xu, W Wei, MH Cheng, BZ Han, XW Chen, DF Su and C Liu. (2017). Autophagy Plays an Important Role in Anti-inflammatory Mechanisms Stimulated by Alpha7 Nicotinic Acetylcholine Receptor. Front Immunol 8:553.
5. Liu M, K Liang, J Zhen, M Zhou, X Wang, Z Wang, X Wei, Y Zhang, Y Sun, Z Zhou, H Su, C Zhang, N Li, C Gao, J Peng and F Yi. (2017). Sirt6 deficiency exacerbates podocyte injury and proteinuria through targeting Notch signaling. Nat Commun 8:413.
6. Chen ZH, WT Wang, W Huang, K Fang, YM Sun, SR Liu, XQ Luo and YQ Chen. (2017). The lncRNA HOTAIRM1 regulates the degradation of PML-RARA oncoprotein and myeloid cell differentiation by enhancing the autophagy pathway. Cell Death Differ 24:212-224.
7. Zhang Y, K Shen, Y Bai, X Lv, R Huang, W Zhang, J Chao, LK Nguyen, J Hua, G Gan, G Hu and H Yao. (2016). Mir143-BBC3 cascade reduces microglial survival via interplay between apoptosis and autophagy: Implications for methamphetamine-mediated neurotoxicity. Autophagy 12:1538-59.
8. Luo T, J Fu, A Xu, B Su, Y Ren, N Li, J Zhu, X Zhao, R Dai, J Cao, B Wang, W Qin, J Jiang, J Li, M Wu, G Feng, Y Chen and H Wang. (2016). PSMD10/gankyrin induces autophagy to promote tumor progression through cytoplasmic interaction with ATG7 and nuclear transactivation of ATG7
9. Hua F, K Li, JJ Yu, XX Lv, J Yan, XW Zhang, W Sun, H Lin, S Shang, F Wang, B Cui, R Mu, B Huang, JD Jiang and ZW Hu. (2015). TRB3 links insulin/IGF to tumour promotion by interacting with p62 and impeding autophagic/proteasomal degradations. Nat Commun 6:7951.
10. Zhang X, Y Yuan, L Jiang, J Zhang, J Gao, Z Shen, Y Zheng, T Deng, H Yan, W Li, WW Hou, J Lu, Y Shen, H Dai, WW Hu, Z Zhang and Z Chen. (2014). Endoplasmic reticulum stress induced by tunicamycin and thapsigargin protects against transient ischemic brain injury: Involvement of PARK2-dependent mitophagy. Autophagy 10:1801-13.








