Background of Coronavirus Disease 2019 (COVID-19, 2019-nCoV)
 Full-text Download
Full-text Download
Author:
Huajun Bai1 Xiaolong Cai1, 2* Xiaoyan Zhang1
1. R&D Center, GeneMedi Co.Ltd., Shanghai, P.R. China (www.genemedi.net)
2. Hanbio Research Center, Hanbio Tech Co. Ltd., Shanghai, P.R. China (www.hanbio.net)
Abstract:
The outbreak of COVID-19, caused by 2019 novel coronavirus (2019-nCoV), has been a global public health threat and caught the worldwide concern. Scientists throughout the world are sparing all efforts to explore strategies for the determination of the 2019-nCoV virus and diagnosis of COVID-19 rapidly. Several assays are developed for COVID-19 test , including RT-PCR, coronavirus antigens-based immunoassays, and CRISPR-based strategies (Cas13a or Cas12a), etc. Different assays have their advantages and drawbacks, and people should choose the most suitable assay according to their demands. Here, we make a brief introduction about these assays and give a simple overview of them, hoping to help doctors and researchers to select the most suitable assay for the Coronavirus Disease 2019 test (COVID-19 test) .
Background of Coronavirus Disease 2019 (COVID-19, 2019-nCoV)
In Dec of 2019, one kind of novel viral pneumonia broke out in Wuhan of China and aroused worldwide concern. This virus was temporarily named as 2019 novel coronavirus (2019-nCoV) by the World Health Organization (WHO) on Jan 7th 2020 [1]. Then, this virus was re-termed as Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) based on its sequence similarity to that of 2002-2003 SARS, and the disease it caused was called Coronavirus Disease 2019 (COVID-19) by WHO on Jan 12th, 2020.
SARS-CoV-2 is 60~200nm in diameter and encapsidates a large single-stranded RNA virus (26-32kb) with many spikes on the virus capsid (Fig. 1A). Several characteristic genes typical for coronaviruses (ORF) are observed in the genomes of SARS-CoV (Fig. 1B), such as spike (S), envelope (E), and nucleocapsid (N). Among them, the receptor-binding domain (RBD) of Spike subunit 1 (S1) is indispensable for the viral infection. To date, the mechanisms about how SARS-CoV transduces human cells have not been completely elucidated yet. Whereas a report shows SARS-CoV 2 infects human with similar processes to SARS by binding to the angiotensin-converting enzyme 2 (ACE2) receptor of target cells [2-4], such as respiratory epithelial cells. Once humans are infected with SARS-CoV-2, they will have the following symptoms: ① having fever and feeling fatigue systematically; ② sneezing, runny nose, sore throat, dry cough, and shortness of breath in the respiratory system; ③ diseased function in kidney; ④ diarrhea in intestines; ⑤ deceased white blood cells.
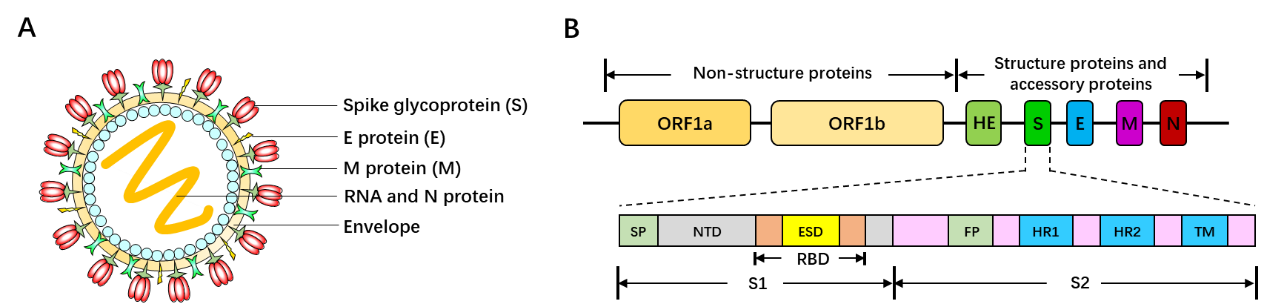
Figure 1. SARS-CoV-2 capsid structure and genome map. (A) Three-dimensional structure diagram of SARS-CoV-2. (B) Genome organization of SARS-CoV-2 [5]. ORF: open reading frame. E: envelope. M: membrane. N: nucleocapsid. HR1: heptad repeat 1. HR2: heptad repeat 2. SP: signal peptide. NTD: N-terminal domain. RBD: receptor binding domain. S: spike. S1: subunit 1. S2: subunit 2. TM: transmembrane domain.
It has been reported that person-to-person transmission from infected COVID-19 patients is really rapidly [5-7]. Although China government is basically in control of COVID-19 pathophoresis by separating patients from normal, forcing citizens to wear masks, and controlling traffic, now the viral pneumonia is globally threatening the health of people all over the world, especially in Europe and North America. Since no specific therapeutic drugs or vaccines are available for patients with COVID-19, it is really necessary and noteworthy to determine whether the patient is infected early and separate the infected patients from the healthy population immediately to avoid the widespread of SARS-CoV-2. To date, there are several strategies for COVID-19 test: ① real-time PCR (RT-PCR) method; ② immunoassay; ③ Crispr-Cas13a (SHERLOCK)-based test.
Reference
1.
L.E.a.V.D.M. Gralinski, Return of the Coronavirus: 2019-nCoV. , Viruses, 2020. 12(2). (2020).
2.
J.L. V. M. Corman, M. Witzenrath, Coronaviruses as the cause of respiratory infections, Internist (Berl) 60, 1136-1145 (2019).
3.
Y. Yang, Q. Lu, M. Liu, Y. Wang, A. Zhang, N. Jalali, N. Dean, I. Longini, M.E. Halloran, B. Xu, X. Zhang, L. Wang, W. Liu, L. Fang, Epidemiological and clinical features of the 2019 novel coronavirus outbreak in China, medRxiv, (2020).
4.
J. Li, S. Li, Y. Cai, Q. Liu, X. Li, Z. Zeng, Y. Chu, F. Zhu, F. Zeng, Epidemiological and Clinical Characteristics of 17 Hospitalized Patients with 2019 Novel Coronavirus Infections Outside Wuhan, China, medRxiv, (2020).
5.
C. Huang, Y. Wang, X. Li, L. Ren, J. Zhao, Y. Hu, L. Zhang, G. Fan, J. Xu, X. Gu, Z. Cheng, T. Yu, J. Xia, Y. Wei, W. Wu, X. Xie, W. Yin, H. Li, M. Liu, Y. Xiao, H. Gao, L. Guo, J. Xie, G. Wang, R. Jiang, Z. Gao, Q. Jin, J. Wang, B. Cao, Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China, The Lancet, 395 (2020) 497-506.
6.
W.-j. Guan, Z.-y. Ni, Y. Hu, W.-h. Liang, C.-q. Ou, J.-x. He, L. Liu, H. Shan, C.-l. Lei, D.S.C. Hui, B. Du, L.-j. Li, G. Zeng, K.-Y. Yuen, R.-c. Chen, C.-l. Tang, T. Wang, P.-y. Chen, J. Xiang, S.-y. Li, J.-l. Wang, Z.-j. Liang, Y.-x. Peng, L. Wei, Y. Liu, Y.-h. Hu, P. Peng, J.-m. Wang, J.-y. Liu, Z. Chen, G. Li, Z.-j. Zheng, S.-q. Qiu, J. Luo, C.-j. Ye, S.-y. Zhu, N.-s. Zhong, Clinical characteristics of 2019 novel coronavirus infection in China, medRxiv, (2020).
7.
N. Chen, M. Zhou, X. Dong, J. Qu, F. Gong, Y. Han, Y. Qiu, J. Wang, Y. Liu, Y. Wei, J.a. Xia, T. Yu, X. Zhang, L. Zhang, Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study, The Lancet, 395 (2020) 507-513.
8.
A. Wu, Y. Peng, B. Huang, X. Ding, X. Wang, P. Niu, J. Meng, Z. Zhu, Z. Zhang, J. Wang, J. Sheng, L. Quan, Z. Xia, W. Tan, G. Cheng, T. Jiang, Genome Composition and Divergence of the Novel Coronavirus (2019-nCoV) Originating in China, Cell Host Microbe, (2020).
9.
R.C. A. R. Fehr, S. Perlman,, Middle East Respiratory Syndrome:Emergence of a Pathogenic Human Coronavirus, Annu Rev Med 68, 387-399 (2017).
10.
X.Y. Ge, Li, J.L., Yang, X.L., Chmura, A.A., Zhu, G.,Epstein, J.H., Mazet, J.K., Hu, B., Zhang, W., Peng,C., et al. , Isolation and characterization of a bat SARS-like coronavirus that uses the ACE2 receptor., Nature 503, 535–538 (2013).
11.
M. Hoffmann, H. Kleine-Weber, N. Krüger, M. Müller, C. Drosten, S. Pöhlmann, The novel coronavirus 2019 (2019-nCoV) uses the SARS-coronavirus receptor ACE2 and the cellular protease TMPRSS2 for entry into target cells, bioRxiv, (2020).
12.
C. Fan, K. Li, Y. Ding, W.L. Lu, J. Wang, ACE2 Expression in Kidney and Testis May Cause Kidney and Testis Damage After 2019-nCoV Infection, medRxiv, (2020).
13.
R. Channappanavar, C. Fett, M. Mack, P.P. Ten Eyck, D.K. Meyerholz, S. Perlman, Sex-Based Differences in Susceptibility to Severe Acute Respiratory Syndrome Coronavirus Infection, The Journal of Immunology, 198 (2017) 4046-4053.
14.
J. Karlberg, D.S. Chong, W.Y. Lai, Do men have a higher case fatality rate of severe acute respiratory syndrome than women do?, Am J Epidemiol, 159 (2004) 229-231.
15.
Z. Li, M. Wu, J. Guo, J. Yao, X. Liao, S. Song, M. Han, J. Li, G. Duan, Y. Zhou, X. Wu, Z. Zhou, T. Wang, M. Hu, X. Chen, Y. Fu, C. Lei, H. Dong, Y. Zhou, H. Jia, X. Chen, J. Yan, Caution on Kidney Dysfunctions of 2019-nCoV Patients, medRxiv, (2020).
16.
W.H. Ding YQ, Shen H, Li ZG, Geng J, Han HX, Cai JJ, Li X, Kang, W.D. W, Lu YD, Wu DH, He L, Yao KT, The clinical pathology of severe acute respiratory syndrome (SARS): a report from China., J Pathol, 2003, 200:282–289 (2003).
17.
Z.L. Lang ZW, Zhang SJ, Meng X, Li JQ, Song CZ, Sun L, Zhou YS, Dwyer DE, A clinicopathological study of three cases of severe acute respiratory syndrome (SARS). , Pathology, 2003, 35:526–531 (2003).
18.
C.P. Chong PY, Ling AE, Franks TJ, Tai DY, Leo YS, Kaw GJ,, C.K. Wansaicheong G, Ean Oon LL, Teo ES, Tan KB, Nakajima, S.T. N, Travis WD, Analysis of deaths during the severe acute respiratory syndrome (SARS) epidemic in Singapore: challenges in determining a SARS diagnosis., Arch Pathol Lab Med, 2004,128:195–204 (2004).
19.
T.W. Chu KH, Tang CS, Lam MF, Lai FM, To KF, Fung KS, Tang HL, Yan WW, Chan HW, Lai TS, Tong KL, Lai KN, Acute renal impairment in coronavirus-associated severe acute respiratory syndrome., Kidney Int, 2005, 67:698–705 (2005).
20.
H.P. Wu VC, Lin WC, Huang JW, Tsai HB, Chen YM, Wu KD, and the SARS Research Group of the National Taiwan, Acute renal failure in SARS patients: more than rhabdomyolysis. , Nephrol Dial Transplant 2004, 19:3180–3182 (2004).
Collection of COVID-19 landscape knowledge base
Viral vector-based vaccine; DNA-based vaccine; RNA based vaccine
- A landscape for vaccine technology against infectious disease, COVID-19 and tumor.
An Insight of comparison between COVID-19 (2019-nCoV disease) and SARS in pathology and pathogenesis
COVID-19 landscape Knowledge Base






