Lentivirus-LC3 production service for Autophagy Flux Detection
Order Information
| Product | Lentivirus Vector Production Scale | Price(In USD) | Qty (Quantity) | Sum(In USD) |
|---|---|---|---|---|
| Lentivirus-mRFP-GFP-LC3 | 1E+8TU | 1880 | ||
| Lentivirus-mRFP-GFP-LC3 | 1E+9TU | 4520 | ||
| Lentivirus-mRFP-GFP-LC3 | 1E+10TU | 15080 | ||
| Lentivirus-GFP-LC3 | 1E+8TU | 1880 | ||
| Lentivirus-GFP-LC3 | 1E+9TU | 4520 | ||
| Lentivirus-GFP-LC3 | 1E+10TU | 15080 | ||
| Shipping Cost: | 760.00 | |||
| Total: | ||||
Introduction to Lentivirus-LC3 production service for Autophagy Flux Detection
Premade LC3 Autophagy Biosensors Products and user manual
Adeno associated virus(AAV) AAV-GFP-LC3 Autophagy BiosensorAAV-mRFP-GFP-LC3 Autophagy Biosensor |
Adenovirus Adv-GFP-LC3 Autophagy BiosensorAdv-mRFP-GFP-LC3 Autophagy Biosensor |
Lentivirus Lv-GFP-LC3 Autophagy BiosensorLv-mRFP-GFP-LC3 Autophagy Biosensor |
LC3 Autophagy Biosensors User Manual
AAV-LC3 User Manual  |
Adenovirus-LC3 User Manual  |
Lentivirus-LC3 User Manual  |
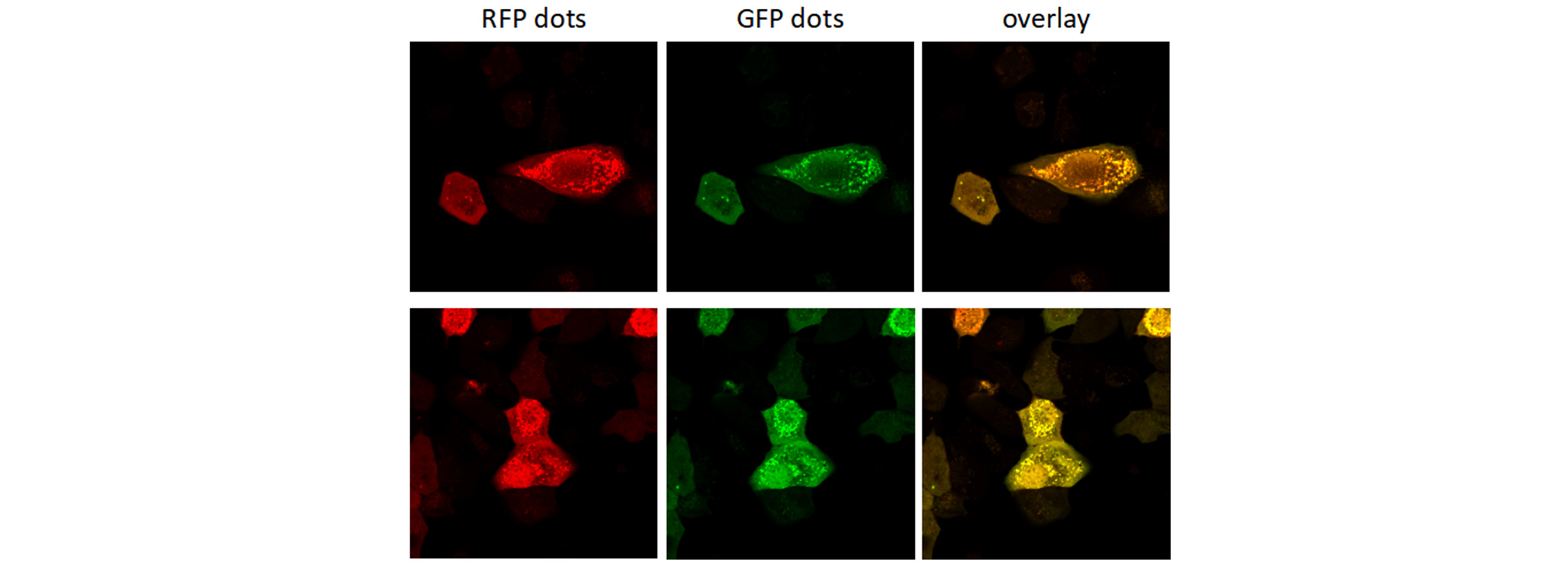
Autophagy is a highly regulated homeostatic degradative process where cells destroy their own components via the lysosomal machinery and recycle them. This process plays both protective and deleterious roles in many diseases, including Alzheimer’s disease, aging, cancer, infection and Crohn’s disease. Elucidating the correlation between autophagy and apoptotic cell death has become the focus of a great deal of research. Members of the LC3 family play a key role in the maturation of the autophagosome. Lysosomal turnover of the autophagosome marker LC3-II reflects starvation-induced autophagic activity, and detecting LC3 by immunoblotting or immunofluorescence has become a reliable method for monitoring autophagy and autophagy-related processes, including autophagic cell death. Genemedi has launched series of lentiviral packaging service of autophagy related biosensors, in which GFP and/or RFP tags are fused at the C-termini of the autophagosome marker LC3, allowing to detect the intensity of autophagy flux in real-time with more accuracy, clarity and intuitiveness. These biosensors provide an enhanced dissection of the maturation of the autophagosome to the autolysosome, which capitalizes on the pH difference between the acidic autolysosome and the neutral autophagosome. The acid-sensitive GFP will be degraded in autolysosome whereas the acid-insensitive RFP will not; Therefore, the change from autophagosome to autolysosome can be visualized by imaging the specific loss of the GFP fluorescence, leaving only red fluorescence. Besides, Genemedi has constructed several other fluorescent biosensors to study the different stages of autophagy flux, making the autophagy study much easier.
Properties
| Lentivirus Production for Autophagy Flux Detection | |
|---|---|
| Quantity/Unit | Vials. |
| Form | Frozen form. |
| Sipping and Storage Guidelines | Shipped by dry ice, stored at -80°C, effective for 1 year. Avoid repeatedly freezing and thawing. |
| Titer | > 1*10^8 TU/ml. |
Advantages
1. Pre-packaged, ready-to-use, fluorescently-tagged with monomeric GFP & RFP.
2. Higher efficiency transfection as compared to traditional chemical-based and other non-viral-based transfection methods. Ability to transfect dividing, non-dividing, and difficult-to-transfect cell types, such as primary cells or stem cells.
3. Non-disruptive towards cellular function.
4. Reveals changing cellular conditions in real time. Enables visualization under different cell/disease states in live cell and in vitro analysis.
Applications and Figures
Quality control description
Our optimized production of lentiviral vector and strict quality control systems supply customers with a high titer of functional recombinant lentiviral vectors. Two methods are employed to determine viral titers: physical titer (VP/mL) and functional titer (TU/mL). Physical titer is calculated by the level of protein, such as p24, or viral nucleic acid. The functional titer, a calculation of the active virus that can infect cells, is much less than the physical titer (100-1000 fold lower). The method we adopted is functional titer, which is an accurate solution for testing virus accurate activity and MOI. The physical titer can only reflect the number of virus particles, but not reflect the true viral activity, which will cause a large error in subsequent infection experiments.
Technical Documents
1. For further information about lentivirus administration and transduction, please see  Lentivirus User Manual.
2. For further information about autophagy study, please see
Lentivirus User Manual.
2. For further information about autophagy study, please see  Lentivirus LC3 Autophagy Flux Detection User Manual.
Lentivirus LC3 Autophagy Flux Detection User Manual.
 Lentivirus User Manual.
2. For further information about autophagy study, please see
Lentivirus User Manual.
2. For further information about autophagy study, please see  Lentivirus LC3 Autophagy Flux Detection User Manual.
Lentivirus LC3 Autophagy Flux Detection User Manual. Frequently Asked Questions(FAQs)
- 1. How can I increase the transduction efficiency?
-
AnswerTry varying particle-to-cell ratio (PPC), incubation volume, temperature and, cell density (if adherent cells are transduced). For adherent cells, we recommend a confluence of about 70%. Following the PPC, adjusting the volume is the next best parameter to change to optimize protein expression.
- 2. What is the difference between LC3A, LC3B and LC3C, or LC3-I and LC3-II?
-
AnswerLC3 is a soluble protein with a molecular mass of ∼17 kDa and is distributed ubiquitously in eukaryotes. It is expressed as the splice variants LC3A, LC3B, and LC3C which display unique tissue distribution. All LC3 isoforms undergo post-translational modifications, especially PE conjugation (lipidation) during autophagy. Upon autophagic signal, the cytosolic form of LC3 (LC3-I) is conjugated to PE to form LC3-PE conjugate (LC3-II), which is recruited to the autophagosomal membranes.
- 3. Can I use the LV-mRFP-GFP-LC3 products to study autophagy in cells in combination with a lentivirus transfection?
-
AnswerLentivirus transfection may actually induce autophagy, skewing the results. Make certain to include appropriate controls and make sure to leave an appropriate time (48-72hrs) post transfection to lower the basal autophagy level.
- 4. Will the GFP signal of mRFP-GFP-LC3 that anchored at autophagosomes and fused with lysosome relight after the cell fixed?
-
AnswerGFP is no longer delighted reversibly once GFP-LC3 signal was destroyed by the acidic environment.
- 5. Can transient expression of RFP-GFP-LC3 differ badly from stable expression?
-
AnswerTransient transfection of GFP-LC3 can leads to the formation of non-autophagic LC3 puncta due to overexpression. Reports shows that GFP-LC3 is aggregate prone protein. We suggest the use of stable cell line for assessment of GFP-LC3 puncta.
- 6. Which software do you usually use for counting the number of LC3 dots in the cell?
-
AnswerImage J does work. For LC3 puncta, we suggest using the "Watershed" plugin from Image J. But proper threshold adjustment is critical. CellProfiler is another choice. First use the top hat filter. Then with the Identify Primary Objects module, manually adjust the Intensity threshold to optimally select the punctas of the right intensity.
- 7. What is a Lentivirus?
-
AnswerLentivirus is a subfamily of the retrovirus family. Lentiviruses can deliver significant amounts of genetic information into host cells and integrate it into the cellular genome. Genetically-engineered lentiviruses are therefore used as one of the most efficient tools of gene delivery. These lentiviruses contain a viral promoter which is used to control the expression of a transgene or shRNA but no virulence genes anymore. Together with several other security modifications this makes them safe to use in the laboratory.
- 8. How does it work?
-
AnswerCo-transfection of packaging plasmids and transfer vector into a packaging cell line allows efficient production of lentiviral particles which are released into the cells supernatant. Viral particles harvested from the cell supernatant can transduce a wide range of both dividing and non-dividing mammalian cell types. Upon infection with lentiviral particles, the single stranded RNA (ssRNA) is reverse-transcribed and the resulting double-stranded DNA (dsDNA) stably integrates into the genome of the host providing for long term transcription of gene of interest or shRNA.
- 9. How long is chloroquine half life when I treat a cell line for an autophagy study?
-
AnswerChloroquine is an attractive drug agent effective for the treatment of not only malaria but also inhibition of autophagy, which is a promising effect for anti-tumor therapy. Half-life time of this drug is approximately 18 hours in vivo due to the degradation by the liver, but the stability of chloroquine in vitro experiments is expected to be longer. It will depend mainly on the cell line your are working on and their metabolic activity. For example HepG2 cells (human hepatocarcinoma cells) have a strong capacity to metabolize drugs. this might not be the case for other cell line of different tissular origin. I would suggest to add chloroquine when you change your growth medium.
- 10. What medium is best to use to induce LC3 puncta in HeLa cells?
-
AnswerWe recommend the media used by Axe et al. (2008) JCB 182: 685-701. The recipe is 140 mM NaCl, 1 mM CaCl2, 1 mM MgCl2, 5 mM glucose, and 20 mM Hepes, pH 7.4, Add 1% BSA and pass through a 20um filter before use.
- 11. What is the best applicable inhibitor of autophagy?
-
AnswerI prefer to use CQ (chloroquine) in all of my researches about autophagy inhibition. The using of CQ is quite easy, since it is easy dissolve in water (unlike 3-MA in DMSO). One time I have been used 3-MA, actually I did not like to work with it. Bafilomycin is another choice for you.
- 12. When to add bafilomycin to study autophagy?
-
Answerit is better to do a time course to be sure. We use at least two times (2 and 4 hours) to starved the cells and we add baf for 2 hours (for 4 hours: the 2 last hours). We then follow LC3-II and P62 by WB. To see the degradation of p62, sommetime 4 hours is better than 2hours.
- 13. Which neuronal cell line should I use for autophagy level?
-
AnswerSHSY5Y and PC12 are two well established neuronal cell lines, human and rat cell lines, respectively. You may also try Neuro2A, it is a mouse cell line.
- 14. How to distinguish selective and non-selective autophagy?
-
AnswerYou can analyze selective mitophagy by interaction/co-localization study of mitochondrial marker protein and the autophagy specific adaptor protein LC3II.
- 15. When to Add the autophagy treament in cell culture models?
-
AnswerFirst, some cells are very sensitive to these inhibitors and you need to optimize your conditions. The second point, if you like to stop autophagy process after inducing it to compare whether it is a cell death or cell survival you may add it just hours (2-3) before treatment. However, many articles prefer to use it as pre-treatment 1 hour before any other treatment.
Reference
1. Wu, J. et al. MicroRNA-30 family members regulate calcium/calcineurin signaling in podocytes. Journal of Clinical Investigation 125, 4091-4106 (2015).
2. Li, F., Li, S. & Cheng, T. TGF-β1 Promotes Osteosarcoma Cell Migration and Invasion Through the miR- 143-Versican Pathway. Cellular Physiology and Biochemistry 34, 2169-2179 (2014).
3. Liu, Z. et al. miR-451a Inhibited Cell Proliferation and Enhanced Tamoxifen Sensitive in Breast Cancer via Macrophage Migration Inhibitory Factor. BioMed Research International 2015, 207684-207684 (2015).
4. Si, L. et al. Smad4 mediated BMP2 signal is essential for the regulation of GATA4 and Nkx2.5 by affecting the histone H3 acetylation in H9c2 cells. Biochemical and Biophysical Research Communications 450, 81-86 (2014).
5. Han, H., Yang, S., Lin, S. G., Xu, C. S. & Han, Z. Effects and mechanism of downregulation of COX‑2 expression by RNA interference on proliferation and apoptosis of human breast cancer MCF‑7 cells. Molecular Medicine Reports 10, 3092-3098 (2014).
6. Zhang, G., Liu, Z., Cui, G., Wang, X. & Yang, Z. MicroRNA-486-5p targeting PIM-1 suppresses cell proliferation in breast cancer cells. Tumor Biology 35, 11137-11145 (2014).
7. Li, G. et al. CYC1 silencing sensitizes osteosarcoma cells to TRAIL-induced apoptosis. Cellular Physiology and Biochemistry 34, 2070-2080 (2014).
8. Mao, J., Lv, Z. & Zhuang, Y. MicroRNA-23a is involved in tumor necrosis factor-α induced apoptosis in mesenchymal stem cells and myocardial infarction. Experimental and Molecular Pathology 97, 23-30 (2014).
9. Liu, X. et al. Role of human pulmonary fibroblast-derived MCP-1 in cell activation and migration in experimental silicosis. Toxicology and Applied Pharmacology 288, 152-160 (2015).
10. Guan, G. et al. CXCR4-targeted near-infrared imaging allows detection of orthotopic and metastatic human osteosarcoma in a mouse model. Scientific Reports 5, 15244-15244 (2015).
11. Zhang, Y. et al. Role of high-mobility group box 1 in methamphetamine-induced activation and migration of astrocytes. Journal of Neuroinflammation 12, 156-156 (2015).
12. Zhu, T. et al. The Role of MCPIP1 in Ischemia/Reperfusion Injury-Induced HUVEC Migration and Apoptosis. Cellular Physiology and Biochemistry 37, 577-591 (2015).
13. Qian, M. et al. P50-associated COX-2 extragenic RNA (PACER) overexpression promotes proliferation and metastasis of osteosarcoma cells by activating COX-2 gene. Tumor Biology 37, 3879-3886 (2016).
14. Wu, N., Song, Y., Pang, L. & Chen, Z. CRCT1 regulated by microRNA-520 g inhibits proliferation and induces apoptosis in esophageal squamous cell cancer. Tumor Biology 37, 8271-8279 (2016).
15. Wang, Y. et al. Overexpression of Hiwi Inhibits the Growth and Migration of Chronic Myeloid Leukemia Cells. Cell Biochemistry and Biophysics 73, 117-124 (2015).
16. Niu, L. et al. RNF43 Inhibits Cancer Cell Proliferation and Could be a Potential Prognostic Factor for Human Gastric Carcinoma. Cellular Physiology and Biochemistry 36, 1835-1846 (2015).
17. Zhang, H. et al. ZC3H12D attenuated inflammation responses by reducing mRNA stability of proinflammatory genes. Molecular Immunology 67, 206-212 (2015).
18. Deng, X. et al. MiR-146b-5p Promotes Metastasis and Induces Epithelial-Mesenchymal Transition in Thyroid Cancer by Targeting ZNRF3. Cellular Physiology and Biochemistry 35, 71-82 (2015).
19. Zhang, B. et al. HSF1 Relieves Amyloid-β-Induced Cardiomyocytes Apoptosis. Cell Biochemistry and Biophysics 72, 579-587 (2015).
20. Hu, Q. et al. Periostin Mediates TGF-β-Induced Epithelial Mesenchymal Transition in Prostate Cancer Cells. Cellular Physiology and Biochemistry 36, 799-809 (2015).
21. Yang, Z. et al. CD49f Acts as an Inflammation Sensor to Regulate Differentiation, Adhesion, and Migration of Human Mesenchymal Stem Cells. Stem Cells 33, 2798-2810 (2015).
22. Wang, X. et al. MCPIP1 Regulates Alveolar Macrophage Apoptosis and Pulmonary Fibroblast Activation After in vitro Exposure to Silica. Toxicological Sciences 151, 126-138 (2016).
23. Gu, S., Ran, S., Liu, B. & Liang, J. miR-152 induces human dental pulp stem cell senescence by inhibiting SIRT7 expression. FEBS Letters 590, 1123-1131 (2016).
24. Jin, F., Qiao, C., Luan, N. & Li, H. Lentivirus-mediated PHLDA2 overexpression inhibits trophoblast proliferation, migration and invasion, and induces apoptosis. International Journal of Molecular Medicine 37, 949-957 (2016).
25. Liu, Z., Song, Y., Wan, L., Zhang, Y. & Zhou, L. Over-expression of miR-451a can enhance the sensitivity of breast cancer cells to tamoxifen by regulating 14-3-3ζ, estrogen receptor α, and autophagy. Life Sciences 149, 104-113 (2016).
26. Tian, Y. et al. MicroRNA-30a promotes chondrogenic differentiation of mesenchymal stem cells through inhibiting Delta-like 4 expression. Life Sciences 148, 220-228 (2016).
27. Xu, S. et al. MicroRNA-33 promotes the replicative senescence of mouse embryonic fibroblasts by suppressing CDK6. Biochemical and Biophysical Research Communications 473, 1064-1070 (2016).
28. Chen, H., Sun, M., Liu, J., Tong, C. & Meng, T. Silencing of Paternally Expressed Gene 10 Inhibits Trophoblast Proliferation and Invasion. PLOS ONE 10 (2015).
29. Deng, Y. et al. Repair of critical-sized bone defects with anti-miR-31-expressing bone marrow stromal stem cells and poly(glycerol sebacate) scaffolds. European Cells & Materials 27, 13-25 (2014).
30. Zheng, Y. & Xu, Z. MicroRNA-22 induces endothelial progenitor cell senescence by targeting AKT3. Cellular Physiology and Biochemistry 34, 1547-1555 (2014).
31. Yang, X. et al. A lentiviral sponge for miRNA-21 diminishes aerobic glycolysis in bladder cancer T24 cells via the PTEN/PI3K/AKT/mTOR axis. Tumor Biology 36, 383-391 (2015).
32. Wang, W. et al. p53/PUMA expression in human pulmonary fibroblasts mediates cell activation and migration in silicosis. Scientific Reports 5, 16900-16900 (2015).
33. Zhang, S. & Qi, Q. MTSS1 suppresses cell migration and invasion by targeting CTTN in glioblastoma. Journal of Neuro-oncology 121, 425-431 (2015).
34. Wang, P. et al. PFDN1, an indicator for colorectal cancer prognosis, enhances tumor cell proliferation and motility through cytoskeletal reorganization. Medical Oncology 32, 264-264 (2015).
35. Gu, S. et al. Human Dental Pulp Stem Cells via the NF-κB Pathway. Cellular Physiology and Biochemistry 36, 1725-1734 (2015).
36. Huang, G. et al. Clinical and therapeutic significance of sirtuin-4 expression in colorectal cancer. Oncology Reports 35, 2801-2810 (2016).
37. Yan, X., Ye, T., Hu, X., Zhao, P. & Wang, X. 58-F, a flavanone from Ophiopogon japonicus, prevents hepatocyte death by decreasing lysosomal membrane permeability. Scientific Reports 6, 27875 (2016).
38. Ding, W., Tong, Y., Zhang, X., Pan, M. & Chen, S. Study of Arsenic Sulfide in Solid Tumor Cells Reveals Regulation of Nuclear Factors of Activated T-cells by PML and p53. Scientific Reports 6, 19793-19793 (2016).
Customer Who View This Product Also Viewed