CRISPR/Cas9 AAV Production Service
Order Information
-
Step 1:
Enter gene information -
Step 2:
AAV vector construction -
Step 3:
AAV packaging -
Step 4:
Custom Order confirmation
| buy | Catalog No. | Vector Type | Vector Name | Promoter | Fluorescent Tag | Lead time(workday) |
| GM-AAVC27342 | Cas9/gRNA Knock-out | AAV-CMV-Sacas9-U6-sgRNA | U6 | null | 10 | |
| GM-AAVC37342 | gRNA Knock-out | pscAAV-U6-BB-chemeric-sgRNA | U6 | null | 10 | |
| GM-AAVC37442 | gRNA Knock-out | pscAAV-U6-BB-sgRNA-CMV-mcherry | U6 | mCherry | 10 | |
| GM-AAVC37142 | gRNA Knock-out | pscAAV-U6-BB-sgRNA-CMV-EGFP | U6 | EGFP | 10 |
| Buy | AAV Vector Production Scale | Final Deliverables | Lead Time(workday) |
|---|
Introduction to CRISPR/Cas9 Adeno Associated Virus (AAV) Production Service
CRISPR/Cas9 premade products
| Cat No. | Products Name | Type of Crispr | Viral vector | Promoter | Fluorescent/Resistance | Tag | Order |
| GMV-Crispr-AAV001 | AAV-CMV-saCas9 | saCas9 | AAV | CMV | Null | HA |  |
| GMV-Crispr-AdV001 | Ad-CMV-spCas9 | spCas9 | Adenovirus | CMV | zsgreen | FLAG |  |
| GMV-Crispr-AdV002 | Adv-CMV-spCas9-zsgreen | spCas9 | Adenovirus | CMV | EGFP | FLAG |  |
| GMV-Crispr-AdV003 | Adv-CMV-spCas9-EGFP | spCas9 | Adenovirus | CMV | Null | FLAG |  |
| GMV-Crispr-AdV004 | Adv-CMV-spCas9-mcherry | spCas9 | Adenovirus | CMV | mcherry | FLAG |  |
| GMV-Crispr-LV001 | Lv-CMV-spCas9-puromycin | spCas9 | lentivirus | CMV | puromycin | FLAG |  |
| GMV-Crispr-LV002 | Lv-CMV-spCas9-zsgreen | spCas9 | lentivirus | CMV | zsgreen | FLAG |  |
| GMV-Crispr-LV003 | Lv-CMV-spCas9-EGFP | spCas9 | lentivirus | CMV | EGFP | FLAG |  |
| GMV-Crispr-LV004 | Lv-CMV-spCas9-mcherry | spCas9 | lentivirus | CMV | mcherry | FLAG |  |
| GMV-Crispr-LV005 | Lv-CBH-spCas9-puromycin | spCas9 | lentivirus | CBH | puromycin | FLAG |  |
| GMV-Crispr-LV006 | Lv-CBH-spCas9-zsgreen | spCas9 | lentivirus | CBH | zsgreen | FLAG |  |
| GMV-Crispr-LV007 | Lv-CBH-spCas9-EGFP | spCas9 | lentivirus | CBH | EGFP | FLAG |  |
| GMV-Crispr-LV008 | Lv-CBH-spCas9-mcherry | spCas9 | lentivirus | CBH | mcherry | FLAG |  |
Custom made CRISPR/Cas9 service
CRISPR/Cas9 User Manual
| Crispr/cas9 mediated Gene knockout  |
AAV Production
CRISPR/Cas9 Knockout System-User Manual  |
Adenovirus
CRISPR/Cas9 Knockout System-User Manual  |
Lentivirus-CRISPR/Cas9 Knockout System-User Manual  |
Gene editing and CRISPR/Cas9
Genemedi has launched a comprehensive AAV packaging service combined with CRISPR/Cas9, the versatile in vitro and
in
vivo genome-editing platform.
Gene editing is a group of technologies to alter the sequence of DNA in the genome. Several approaches to genome
editing have been developed, including ZFN, TALEN and CRISPR/Cas9 technologies. Among them, the CRISPR/Cas9
system
is much faster, cheaper, more accurate, and more efficient than other existing genome editing methods. Cas9, an
RNA-guided endonuclease derived from the type II CRISPR-Cas bacterial adaptive immune system, has emerged as a
versatile genome-editing platform.
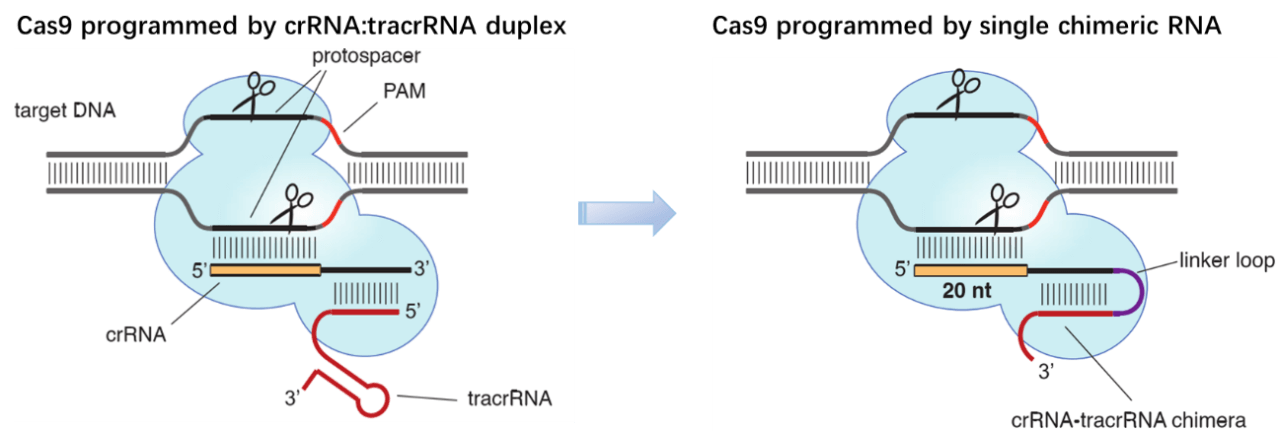
Figure 1. Cas9 can be programmed using a single engineered RNA molecule covering tracrRNA and crRNA features. Left, Cas9 is directed by a dual RNA structure formed by activating tracrRNA and targeting crRNA to cleave site-specifically–targeted dsDNA in type II CRISPR/Cas systems. Right, a chimeric RNA generated through fusion of crRNA to tracrRNA [1].
Adeno associated virus (AAV)Adeno associated virus (AAV) is one kind of human parvovirus. Recombinant AAV vectors can efficiently transfect various cell types, including dividing and quiescent cells, and induce persistent gene expression in vivo without integrating into host genome and causing any disease. These features make AAV an attractive candidate in the application of gene delivery for gene therapy and human disease model establishment. To date, AAV has been proved as the most excellent gene therapy vector. Over 204 clinical trials have been carried out using AAV vectors for gene delivery, and promising gene therapy outcomes have been achieved from clinical trials for a great number of diseases.
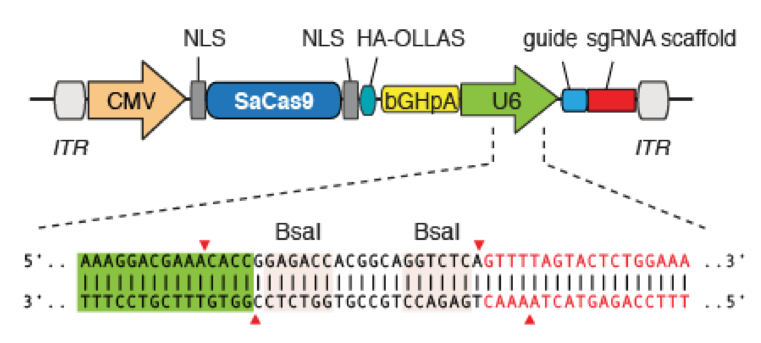
Genemedi AAV-SaCas9 production serviceGenemedi has launched a comprehensive AAV packaging service combined with CRISPR/Cas9, the versatile genome-editing platform. However, the restrictive cargo size (~4.5kb) of AAV presents an obstacle for AAV packaging the conventionally used SpCas9 and its single guide RNA (sgRNA) in a single vector. In 2015, Kleinstiver et al discovered one smaller-size Cas9 orthologues, Staphylococcus aureus Cas9 (SaCas9), which is 25% smaller than SpCas9, efficient and broadly targeting [2]. In combination with AAV Vector System, Genemedi has further developed AAV-SaCas9 with different tropisms and serotypes to achieve in vivo gene knockout with high efficiency, which is one of the top advanced in vivo gene knockout technology in the world.
Properties
| Genemedi AAV SaCas9 Particles | |
|---|---|
| Quantity/Unit | Vials |
| Form | Frozen form |
| Suitable Types of Infection | In vivo infection in animals |
| Sipping and Storage Guidelines | Shipped by dry ice, stored at -80 ° C, effective for 1 year. Avoid repeatedly freezing and thawing. |
| Titer | > 1*10^12v.g/ml |
Advantages
1. Genome editing system of somatic tissues in postnatal animals.
2. Utilizing Staphylococcus aureus Cas9 (SaCas9). SaCas9, is 25% smaller than SpCas9, thus can be packaged into AAV for in vivo genome editing.
3. Safety. The wild type Adeno Associated Virus (AAV) has not currently been known to cause disease, and further psecurity of recombinant AAV is ensured after removal of most AAV genome elements.
4. Low immunogenicity. AAV causes a very mild immune response, lending further support to its apparent lack of pathogenicity.
5. Broad range of host and specificity targeting. AAV has the ability to infect both dividing and quiescent cells, allowing genetic material to be delivered to a highly diverse range of cell types. More than 12 AAV serotypes and a variety of capsid engineered AAV vectors can be selected according to their tissue tropisms.
6. Broad range of host and specificity targeting. Stable physical properties. AAV is still alive at 60 ℃ and resistant to chloroform.
7. Stable physical properties. AAV is still alive at 60 ℃ and resistant to chloroform.
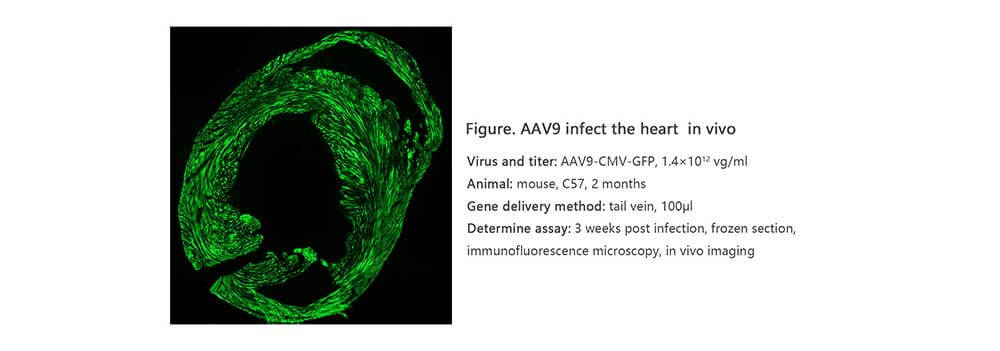
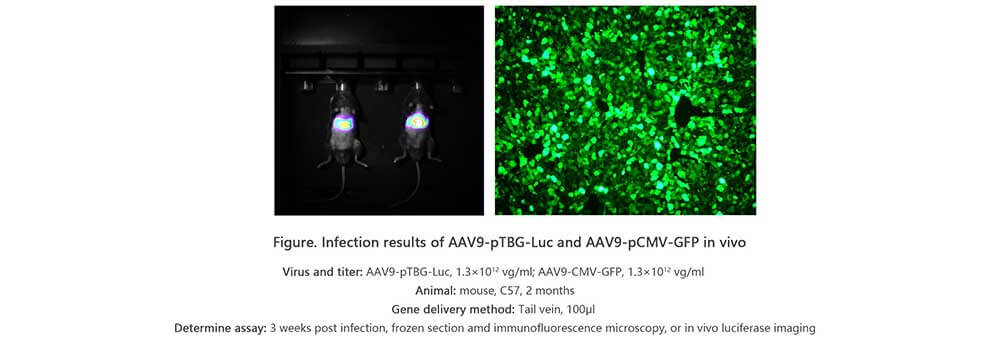
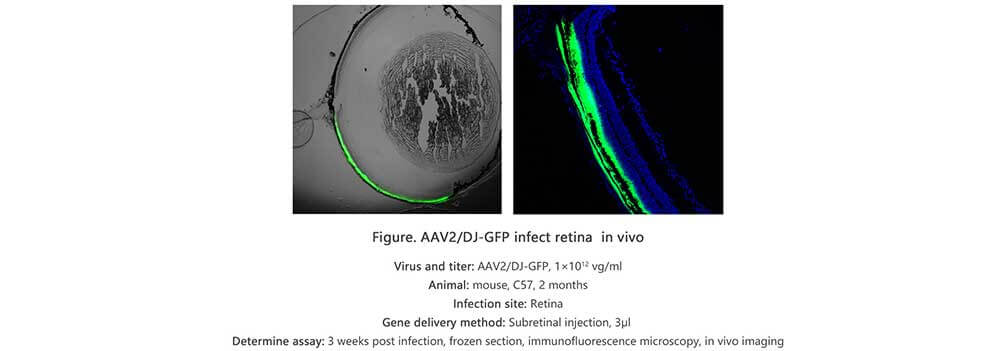
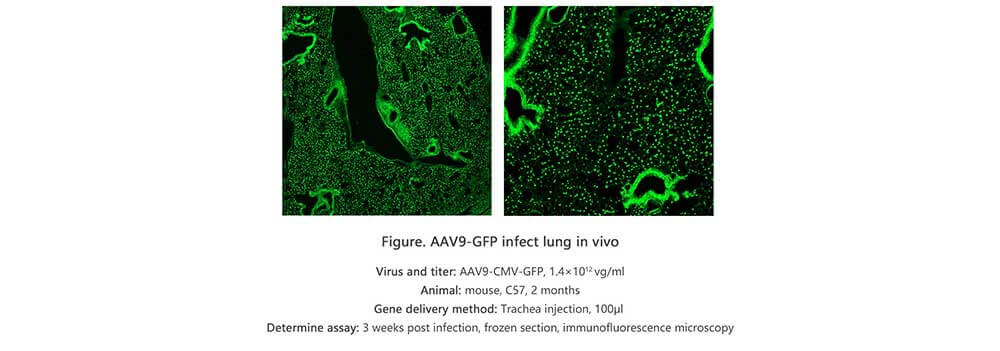
Applications and Figures
Quality control description
AAV capsid contains VR1 82kDa, VR2 72kDa and VR3 62kDa, which can be detected using the method of polyacrylamide gel electrophoresis (PAGE) followed by silver staining or Coomassie blue staining. Pure AAV should display only three major protein bands, such as shown in Fig 3.
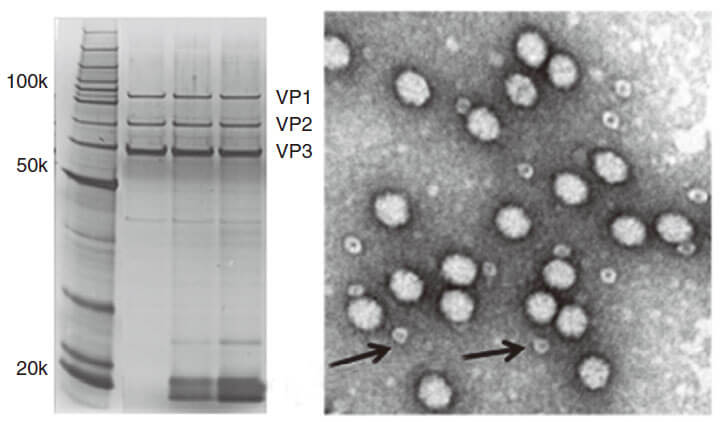 Figure 3. PAGE for quality testing example of AAV by Genemedi. Only 3 protein bands can be seen in the purified AAV virus samples.
Figure 3. PAGE for quality testing example of AAV by Genemedi. Only 3 protein bands can be seen in the purified AAV virus samples.Technical Documents
1. For further information about AAV administration and transduction results, please see the  AAV(adeno associated virus) User Manual.
AAV(adeno associated virus) User Manual.
2. For more information about how to design AAV-SaCas9 system, please refer to the  AAV-SaCas9 User Manual on the Genemedi website.
AAV-SaCas9 User Manual on the Genemedi website.
Frequently Asked Questions(FAQs)
- 1. How can I choose the optimal AAV serotypes for my in vivo study?
-
Answer
Genemedi has launched a comprehensive AAV production service. More than 12 AAV serotypes and a variety of capsid engineered AAV vectors are available for targeting different tissues and organs. Different AAV serotypes have different tissue tropism in vivo, the common serotypes and their tropism are listed in the above table 1. Therefore, you can select the most suitable AAV serotype for your study according to this table. For example, if you’d like to transfect your target gene into mouse ears, the AAV-Anc80 will be your best choice.
For your optimal AAV serotypes screening, Genemedi can also customize an AAV serotype testing kit with recommended serotypes of your choice. - 2. How much DNA do I need to provide for Custom AAV without DNA amplification?
-
Answer
You will need to provide purified plasmid DNA at a concentration of 0.5ug/ul or more.
50 ug DNA needed for Custom AAV production service without DNA amplification (10^9 GC/ml).
200 ug DNA needed for Custom AAV production service without DNA amplification (10^12 GC/ml).
300 ug DNA needed for Custom AAV production service without DNA amplification (10^13 GC/ml). - 3. What's the difference of AAV titer unit between GC/ml and vg/ml?
-
Answer
The AAV titer unit GC/ml and vg/ml can be used interchangeably, based on qPCR method.
- 4. Why can’t I see the expression of GFP after AAV infection in my cell line?
-
Answer
AAV serotypes selection is an important parameter which may affect the transduction ability of AAV particles. Thus, it is necessary to determine which serotype works best for your cell line. For instance, serotype 5 limits its transduction ability on most cell types. For detailed information, you can refer to our technical sheet "AAV - General Guideline to Serotype Selection".
- 5. Does Genemedi’s AAV preparations belong to in-vivo grade?
-
Answer
Extensive purification steps guarantee the high quality of our viral particles, which are ready to be administrated for in-vivo models. For detailed references, please see the in-vivo infection data tested by another independent lab.
- 6. What is the QC (quality control) method for testing AAV in Genemedi?
-
Answer
We provide qPCR-based titer as the primary method to determine whether the packaging/purification process is successful or not. If your virus has GFP or RFP reporter, we also perform virus infectivity testing in HEK293T cell line.
- 7. Can I utilize different serotypes of AAV virus in the same equipment to keep the infected cells?
-
Answer
There is no problem with using different serotypes in the same equipment, as long as the handler takes the basic precautions to avoid cross-contamination.
- 8. What is the difference between brain localization of gene expression after injection of AAV or lentiviral vectors?
-
Answer
Lentiviral particles don't spread well after stereotaxic injection into brain because the particles are relatively large (90nm~100nm). In contrast, AAV particles spread more readily due to smaller size (20nm~100nm). As mentioned, it has been stated that some AAV serotypes spread better than others in brain, for example, AAV5 is reported to spread exceptionally well when injected into striatum. Lastly, pretreatment with mannitol to your animal about 15 min ahead of viral injection has been reported to aid in spread of viral particles in the brain.
- 9. Do AAVs tranduce axons passing through a region of interest?
-
Answer
AAVs are able to infect axon terminals and produce retrograde transport (towards the cell body). The slightly longer answer is that this process is highly biased based on the serotype you are using. There are a number of papers reporting AAV6 (and maybe AAV6.2) is the main serotype to produce retrograde transport. There are however reports of many other serotypes producing retrograde transport with a much smaller rate.
- 10. Is it still possible to get the same mRNA accumulation in RT PCR after CRISPR/Cas9 induced gene deletion?
-
Answer
Yes. The transcript will continue to be transcribed as nothing has changed with respect to the promoter of the gene. Besides the indel you introduced, the transcript will be unchanged and your primers (presumably not overlapping indel) will detect the transcript. You'll want to use Western blot to observe the effect of the indel.
- 11. I am trying to design gRNA for TLR4 gene CRISPR cas9 genome editing. TLR4 has multiple transcript variants. Which one should I choose?
-
Answer
You should use a gRNA which target as many transcript variants as possible. To make it clear, target those exon(s) which can be found in the most variants of TLR4. As I see exon 1 and exon 4 are the best choices to target TLR4.
- 12. Which sequencing depth should I use for my CRISPR-Cas9 screen?
-
Answer
PCR amplification and Sanger sequencing are the main CRISPR KO screening methods.
- 13. Does the CRISPR/Cas9 system work for non-sequenced organisms?
-
Answer
In theory it should be able to work with any organism. However, there may be some modifications that you will need to work out. You may need to codon optimize the Cas9 gene for your organism and confirm if there are many rare codons used in your organism. Next, you'll need to find a good promoter in your organism that you can use to make your crRNA. Maybe you'll need to add a nucleus localization signal to the Cas9 to make sure it stays in the nucleus, for Cas9 won't work if it is in the cytoplasm. Another issue you need to determine is what protospacer to use. If your organism is similar to one that is sequenced, then this may work, otherwise if you don't know the DNA sequence of your gene, it will be impossible to design a good protospacer against it. Lastly, you'll need a way to transfer Cas9 and crRNA into your organism, which should not hurt your organism.
Reference
1.
Kleinstiver BP, MS Prew, SQ Tsai, VV Topkar, NT Nguyen, Z Zheng, AP Gonzales, Z Li, RT Peterson, JR Yeh, MJ Aryee and JK Joung. (2015). Engineered CRISPR-Cas9 nucleases with altered PAM specificities. Nature 523:481-5.
2.
Ran FA, L Cong, WX Yan, DA Scott, JS Gootenberg, AJ Kriz, B Zetsche, O Shalem, X Wu, KS Makarova, EV Koonin, PA Sharp and F Zhang. (2015). In vivo genome editing using Staphylococcus aureus Cas9. Nature 520:186-91.
3.
Jinek M, K Chylinski, I Fonfara, M Hauer, JA Doudna and E Charpentier. (2012). A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337:816-21.
4.
Kleinstiver BP, MS Prew, SQ Tsai, VV Topkar, NT Nguyen, Z Zheng, AP Gonzales, Z Li, RT Peterson, JR Yeh, MJ Aryee and JK Joung. (2015). Engineered CRISPR-Cas9 nucleases with altered PAM specificities. Nature 523:481-5.
5.
Ran FA, L Cong, WX Yan, DA Scott, JS Gootenberg, AJ Kriz, B Zetsche, O Shalem, X Wu, KS Makarova, EV Koonin, PA Sharp and F Zhang. (2015). In vivo genome editing using Staphylococcus aureus Cas9. Nature 520:186-91.