2019 nCoV (SARS CoV2 coronavirus) Ad-Spike (S1+S2) whole mutation variant of SARS-CoV-2(2019nCoV) 501Y.V2 lineage(B.1.351) Pre-made adenovirus vector
Cat No.:GMV-V-2019nCoV-112
Order information
| Catalog No. | Package | Price(In USD) | Qty (Quantity) | Sum(In USD) |
|---|---|---|---|---|
| GMV-V-2019nCoV-112 | In vitro grade: 1E+10PFU (1E+10pfu/ml×1ml) | 3790 | ||
| GMV-V-2019nCoV-112 | In vitro grade: 5E+10PFU (1E+10pfu/ml×5ml) | 5390 | ||
| GMV-V-2019nCoV-112 | In vitro grade: 1E+11PFU (1E+10pfu/ml×10ml) | 9890 | ||
| GMV-V-2019nCoV-112 | Large size | Inquiry. | ||
| GMV-V-2019nCoV-112 | In vivo grade: 1E+11PFU (1E+11pfu/ml×1ml) | Inquiry. | ||
| Shipping Cost: | 760.00 | |||
| Total: | ||||
Description
| 2019 nCoV related Gene | Spike (S1+S2) whole mutation variant of SARS-CoV-2(2019nCoV) 501Y.V2 lineage(B.1.351) |
| Species Host | SARS-COV-2 501Y.V2 lineage(B.1.351) |
| Vector | Pre-made adenovirus |
| Reporter | null |
| Antibiotic in Mammalian cell | null |
| Length | 3822 |
| Promoter | CMV |
| Tag | C-3FLAG |
| Codon Optimized | Codon Optimized for mamamlian |
| Gene Accession Number | QHD43416.1 |
| Application | Adenovirus-based 2019nCoV vaccine; Adenovirus production; Mammalian cells expression |
Spike mutant variant of SARS-COV-2 (2019nCOV) 501Y.V2 lineage(B.1.351) spread in South Africa
|
Background Reading: Information and products collection of SARS-COV-2 (2019nCOV) South Africa 501Y.V2 lineage |
Mutations of Spike protein in SARS-COV-2 (2019nCOV) 501Y.V2 lineage(B.1.351) spread in South Africa
Three of the spike mutations are at key residues in the RBD (N501Y, E484K and K417N), three are in the N-terminal domain (L18F, D80A and D215G) and one is in loop 2 (A701V). Deletion of three amino acids at L242-244L is disputed with a L242H mutation1.| Spike Mutation in SARS-COV-2 (2019nCOV) 501Y.V2 lineage(B.1.351) |
Spike-S1 Subunit N Terminal |
L18F | |
| D8OA | |||
| D215G | |||
| Spike-S1 Subunit | L242-244 deletion | disputed with a L242H mutation | |
| Spike-S1 Subunit | R246I | ||
| Spike-RBD | K417N | Important residues mutation | |
| E484K | |||
| N501Y | |||
| Spike-S1 Subunit | D614G | A popular mutation in different new SARS-CoV-2 lineage | |
| Spike-S2 Subunit | A701V |
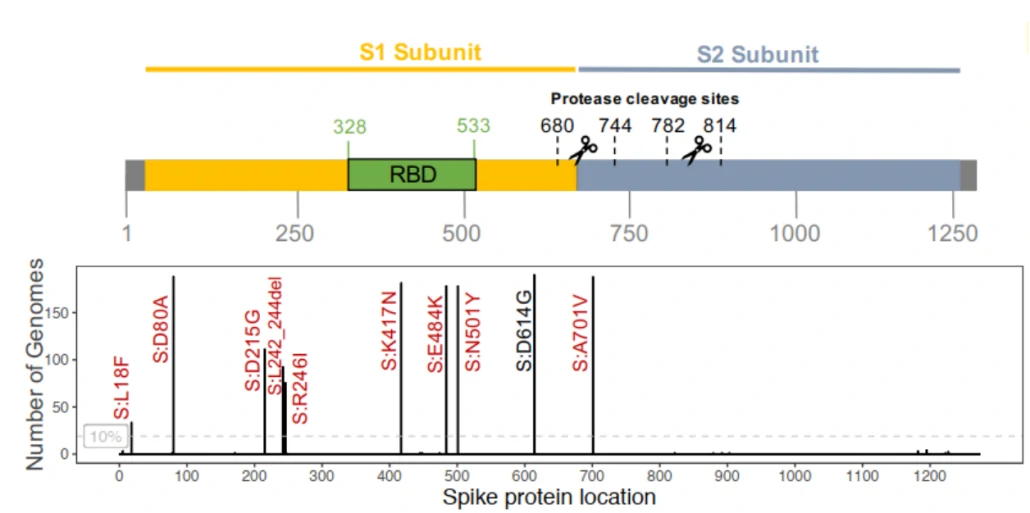
Figure. Amino acid changes in the spike region of the 190 S501Y.V2 genomes1
GeneMedi codon-optimized spike mammalian expression vector for SARS-COV-2 (2019nCOV) S501Y.V2 lineage(B.1.351)
GeneMedi pseudotype virus (pseudovirus) of SARS-COV-2 (2019nCOV) 501Y.V2 lineage(B.1.351)
Reference1 Houriiyah Tegally, E. W., Marta Giovanetti, Arash Iranzadeh, Vagner Fonseca, Jennifer Giandhari, Deelan Doolabh, Sureshnee Pillay, Emmanuel James San, Nokukhanya Msomi, Koleka Mlisana, Anne von Gottberg, Sibongile Walaza, Mushal Allam, Arshad Ismail, Thabo Mohale, Allison J Glass, Susan Engelbrecht, Gert Van Zyl, Wolfgang Preiser, Francesco Petruccione, Alex Sigal, Diana Hardie, Gert Marais, Marvin Hsiao, Stephen Korsman, Mary-Ann Davies, Lynn Tyers, Innocent Mudau, Denis York, Caroline Maslo, Dominique Goedhals, Shareef Abrahams, Oluwakemi Laguda-Akingba, Arghavan Alisoltani-Dehkordi, Adam Godzik, Constantinos Kurt Wibmer, Bryan Trevor Sewell, José Lourenço, Luiz Carlos Junior Alcantara, Sergei L Kosakovsky Pond, Steven Weaver, Darren Martin, Richard J Lessells, Jinal N Bhiman, Carolyn Williamson, View ORCID ProfileTulio de Oliveira. Emergence and rapid spread of a new severe acute respiratory syndrome-related coronavirus 2 (SARS-CoV-2) lineage with multiple spike mutations in South Africa. medRxiv preprint, doi:https://doi.org/10.1101/2020.12.21.20248640 (2020).

<







