Pre-made gene ORF plasmids for 2019 nCoV (SARS2 coronavirus, COVID-19)
N protein, S protein(Spike), RBD, E protein(Envelope), M protein, ORF1ab/3a/6/7/8/10
Content index
About Pre-made gene ORF plasmids for 2019 nCoV (SARS2 coronavirus, COVID-19)
COVID-19, a novel pathogenic coronavirus (2019-nCoV or SARS-CoV2) induced disease, emerged in China and spread globally rapidly.
GeneMedi offers pre-made gene ORF plasmids for 2019 nCoV (SARS2 coronavirus, COVID-19) as shown below: N protein(Nucleocapsid), S protein(Spike, full lenth), S1 protein(Spike), S protein RBDS protein (Spike receptor binding domain), Spike mutation (mutation of D614G, S943P, V367F, G476S, V483A, H49Y, Q239K, A831V, P1263L, D839Y/N/E, D839Y, D839N, D839E, P2-mutated Spike protein trimer variant ( P2-mutant, S1/S2 cleavage site(furin cleavage sequence)-mutant, trimerization modified), Spike(S1+S2) - B.1.1.7 lineage, Spike(S1+S2)-N501Y mutation, Spike (S1+S2)-HV 69-70 Deletion mutation(ΔH69/ΔV70) ), E protein(Envelope), M protein, ORF1ab/3a/6/7/8/10.
The pre-made gene ORF vector of 2019 nCoV (SARS2 coronavirus) can be used for mammalian expression or E.coli expression dependent on the CMV promoter or T7 protomer.
GeneMedi also provides pre-made the lentivirus, adenovirus and AAV vector for the gene ORF plasmids of 2019 nCoV (SARS2 coronavirus).
GeneMedi has submitted a preprints review about COVID-19 and 2019-nCoV.
Click here for the full-text reading:
Viral vector-based vaccine; DNA-based vaccine; RNA based vaccine
- A landscape for vaccine technology against infectious disease, COVID-19 and tumor.
An Insight of comparison between COVID-19 (2019-nCoV disease) and SARS in pathology and pathogenesis
Landscape Coronavirus Disease 2019 test (COVID-19 test) in vitro -- A comparison of PCR vs Immunoassay vs Crispr-Based test
In the mission of helping global scientists for the development of diagnosis and drug against 2019 nCoV(SARS2 coronavirus), GeneMedi scientists will keep moving.
Pre-made gene ORF plasmids for 2019 nCoV (SARS CoV-2 coronavirus, COVID-19) Spike protein(S protein):
Spike full length, Spike RBD,Spike mutation(D614G mutataion,S943P,V367F, G476S, V483A, H49Y, Q239K, A831V, P1263L, D839Y/N/E,D839Y, D839N, D839E,
P2-mutated Spike protein trimer variant ( P2-mutant,S1/S2 cleavage site(furin cleavage sequence)-mutant,trimerization modified), Spike(S1+S2) - B.1.1.7 lineage,
Spike(S1+S2)-N501Y mutation, Spike(S1+S2)-HV 69-70 Deletion mutation(ΔH69/ΔV70)
2019 nCoV (SARS2 coronavirus) gene and vectors
| Cat No. | 2019 nCoV related Gene |
Gene &Vector description of 2019 nCoV | Vector | Reporter | Tag | codon Optimized |
Order |
| GMV-V-2019nCoV-015 | ORF8 | pGM-2019nCoV-ORF8 | pcDNA3.1(+) | null | No tag | No |  |
| GMV-V-2019nCoV-024 | ORF8 | pGM-H2019nCoV-ORF8 | pCDNA3.1(+) | null | No tag | Codon Optimized for mammalian |  |
| GMV-V-2019nCoV-033 | ORF8 | pGM-E2019nCoV-ORF8 | pET21a | null | C-His | Codon Optimized for E.coli |  |
| GMV-V-2019nCoV-014 | ORF7 | pGM-2019nCoV-ORF7a | pcDNA3.1(+) | null | No tag | No |  |
| GMV-V-2019nCoV-023 | ORF7 | pGM-H2019nCoV-ORF7a | pCDNA3.1(+) | null | No tag | Codon Optimized for mammalian |  |
| GMV-V-2019nCoV-032 | ORF7 | pGM-E2019nCoV-ORF7a | pET21a | null | C-His | Codon Optimized for E.coli |  |
| GMV-V-2019nCoV-013 | ORF6 | pGM-2019nCoV-ORF6 | pcDNA3.1(+) | null | No tag | No |  |
| GMV-V-2019nCoV-022 | ORF6 | pGM-H2019nCoV-ORF6 | pCDNA3.1(+) | null | No tag | Codon Optimized for mammalian |  |
| GMV-V-2019nCoV-031 | ORF6 | pGM-E2019nCoV-ORF6 | pET21a | null | C-His | Codon Optimized for E.coli |  |
| GMV-V-2019nCoV-012 | ORF3a | pGM-2019nCoV-ORF3a | pcDNA3.1(+) | null | No tag | No |  |
| GMV-V-2019nCoV-021 | ORF3a | pGM-H2019nCoV-ORF3a | pCDNA3.1(+) | null | No tag | Codon Optimized for mammalian |  |
| GMV-V-2019nCoV-030 | ORF3a | pGM-E2019nCoV-ORF3a | pET21a | null | C-His | Codon Optimized for E.coli |  |
| GMV-V-2019nCoV-016 | ORF10 | pGM-2019nCoV-ORF10 | pcDNA3.1(+) | null | No tag | No |  |
| GMV-V-2019nCoV-025 | ORF10 | pGM-H2019nCoV-ORF10 | pCDNA3.1(+) | null | No tag | Codon Optimized or mammalian |  |
| GMV-V-2019nCoV-034 | ORF10 | pGM-E2019nCoV-ORF10 | pET21a | null | C-His | Codon Optimized for E.coli |  |
| GMV-V-2019nCoV-011 | N protein | pGM-2019nCoV-Nucleocapsid Protein (N protein) | pcDNA3.1(+) | null | No tag | No |  |
| GMV-V-2019nCoV-020 | N protein | pGM-2019nCoV-Nucleocapsid Protein (N protein) | pCDNA3.1(+) | null | No tag | Codon Optimized for mammalian |  |
| GMV-V-2019nCoV-029 | N protein | pGM-2019nCoV-Nucleocapsid Protein (N protein) | pET21a | null | C-His | Codon Optimized for E.coli |  |
| GMV-V-2019nCoV-010 | M protein | pGM-2019nCoV-Membrane protein (M protein) | pcDNA3.1(+) | null | No tag | No |  |
| GMV-V-2019nCoV-019 | M protein | pGM-2019nCoV-Membrane protein (M protein) | pCDNA3.1(+) | null | No tag | Codon Optimized for mammalian |  |
| GMV-V-2019nCoV-028 | M protein | pGM-2019nCoV-Membrane protein (M protein) | pET21a | null | C-His | Codon Optimized for E.coli |  |
| GMV-V-2019nCoV-009 | E protein | pGM-2019nCoV-Envelope Protein (E protein) | pcDNA3.1(+) | null | No tag | No |  |
| GMV-V-2019nCoV-018 | E protein | pGM-2019nCoV-Envelope Protein (E protein) | pCDNA3.1(+) | null | No tag | Codon Optimized for mammalian |  |
| GMV-V-2019nCoV-027 | E protein | pGM-2019nCoV-Envelope Protein (E protein) | pET21a | null | C-His | Codon Optimized for E.coli |  |
| GMV-V-2019nCoV-045 | TMPRSS2 | pGMLv-hTMPRSS2(C-3FLAG) | Lentiviral vector | Zsgreen | C-3FLAG | No |  |
| GMV-V-2019nCoV-041 | ACE2 | pGMLV-hACE2(C-3FLAG) | Lentiviral vector | Zsgreen | C-3FLAG | No |  |
| GMV-V-2019nCoV-042 | ACE2 | pAD-hACE2(C-3FLAG) | Pre-made Adenovirus | EGFP | C-3FLAG | No |  |
| GMV-V-2019nCoV-046 | TMPRSS2 | pGMLv-mtmprss2(C-3FLAG) | Lentiviral vector | Zsgreen | C-3FLAG | No |  |
| GMV-V-2019nCoV-043 | ACE2 | pAD-mACE2(C-3FLAG) | Pre-made Adenovirus | EGFP | C-3FLAG | No |  |
| GMV-V-2019nCoV-044 | ACE2 | pGMLV-mACE2(C-3FLAG) | Lentiviral vector | Zsgreen | C-3FLAG | No |  |
| GMV-V-2019nCoV-068 | Nsp1 | pGM-2019nCoV-Nsp1 | Lentiviral vector | null | 2×Strep | Codon Optimized for mammalian |  |
| GMV-V-2019nCoV-069 | Nsp1 | pGM-2019nCoV-Nsp1 | pCDNA6B | null | FLAG | No |  |
| GMV-V-2019nCoV-070 | Nsp2 | pGM-2019nCoV-Nsp2 | Lentiviral vector | null | 2×Strep | Codon Optimized for mammalian |  |
| GMV-V-2019nCoV-071 | Nsp2 | pGM-2019nCoV-Nsp2 | pCDNA6B | null | FLAG | No |  |
| GMV-V-2019nCoV-072 | Nsp3C | pGM-2019nCoV-Nsp3C | pCDNA6B | null | FLAG | No |  |
| GMV-V-2019nCoV-073 | Nsp3N | pGM-2019nCoV-Nsp3N | pCDNA6B | null | FLAG | No |  |
| GMV-V-2019nCoV-074 | Nsp4 | pGM-2019nCoV-Nsp4 | Lentiviral vector | null | 2×Strep | Codon Optimized for mammalian |  |
| GMV-V-2019nCoV-075 | Nsp4 | pGM-2019nCoV-Nsp4 | pCDNA6B | null | FLAG | No |  |
| GMV-V-2019nCoV-076 | Nsp5(Mpro,3CLpro) | pGM-2019nCoV-Nsp5(Mpro,3CLpro) | Lentiviral vector | null | 2×Strep | Codon Optimized for mammalian |  |
| GMV-V-2019nCoV-077 | Nsp5(Mpro,3CLpro) | pGM-2019nCoV-Nsp5(Mpro,3CLpro) | pCDNA6B | null | FLAG | No |  |
| GMV-V-2019nCoV-078 | Nsp6 | pGM-2019nCoV-Nsp6 | Lentiviral vector | null | 2×Strep | Codon Optimized for mammalian |  |
| GMV-V-2019nCoV-079 | Nsp6 | pGM-2019nCoV-Nsp6 | pCDNA6B | null | FLAG | No |  |
| GMV-V-2019nCoV-080 | Nsp7 | pGM-2019nCoV-Nsp7 | Lentiviral vector | null | 2×Strep | Codon Optimized for mammalian |  |
| GMV-V-2019nCoV-081 | Nsp7 | pGM-2019nCoV-Nsp7 | pCDNA6B | null | FLAG | No |  |
| GMV-V-2019nCoV-082 | Nsp8 | pGM-2019nCoV-Nsp8 | Lentiviral vector | null | 2×Strep | Codon Optimized for mammalian |  |
| GMV-V-2019nCoV-083 | Nsp8 | pGM-2019nCoV-Nsp8 | pCDNA6B | null | FLAG | No |  |
| GMV-V-2019nCoV-084 | Nsp9 | pGM-2019nCoV-Nsp9 | Lentiviral vector | null | 2×Strep | Codon Optimized for mammalian |  |
| GMV-V-2019nCoV-085 | Nsp9 | pGM-2019nCoV-Nsp9 | pCDNA6B | null | FLAG | No |  |
| GMV-V-2019nCoV-086 | Nsp10 | pGM-2019nCoV-Nsp10 | Lentiviral vector | null | 2×Strep | Codon Optimized for mammalian |  |
| GMV-V-2019nCoV-087 | Nsp10 | pGM-2019nCoV-Nsp10 | pCDNA6B | null | FLAG | No |  |
| GMV-V-2019nCoV-088 | Nsp11 | pGM-2019nCoV-Nsp11 | Lentiviral vector | null | 2×Strep | Codon Optimized for mammalian |  |
| GMV-V-2019nCoV-089 | Nsp11 | pGM-2019nCoV-Nsp11 | pCDNA6B | null | FLAG | No |  |
| GMV-V-2019nCoV-090 | Nsp12(RdRP) | pGM-2019nCoV-Nsp12(RdRP) | Lentiviral vector | null | 2×Strep | Codon Optimized for mammalian |  |
| GMV-V-2019nCoV-091 | Nsp12(RdRP) | pGM-2019nCoV-Nsp12(RdRP) | pCDNA6B | null | FLAG | No |  |
| GMV-V-2019nCoV-092 | Nsp13 | pGM-2019nCoV-Nsp13 | Lentiviral vector | null | 2×Strep | Codon Optimized for mammalian |  |
| GMV-V-2019nCoV-093 | Nsp13 | pGM-2019nCoV-Nsp13 | pCDNA6B | null | FLAG | No |  |
| GMV-V-2019nCoV-094 | Nsp14 | pGM-2019nCoV-Nsp14 | Lentiviral vector | null | 2×Strep | Codon Optimized for mammalian |  |
| GMV-V-2019nCoV-095 | Nsp14 | pGM-2019nCoV-Nsp14 | pCDNA6B | null | FLAG | No |  |
| GMV-V-2019nCoV-096 | Nsp15 | pGM-2019nCoV-Nsp15 | Lentiviral vector | null | 2×Strep | Codon Optimized for mammalian |  |
| GMV-V-2019nCoV-097 | Nsp15 | pGM-2019nCoV-Nsp15 | pCDNA6B | null | FLAG | No |  |
| GMV-V-2019nCoV-098 | Nsp16 | pGM-2019nCoV-Nsp16 | pCDNA6B | null | FLAG | No |  |
Validation of SARS-CoV-2 (2019nCoV) codon-optimized (CO) Spike overexpression in HEK293 cell line by GeneMedi's adenoviral and lentiviral vector
 Post Download
Post Download
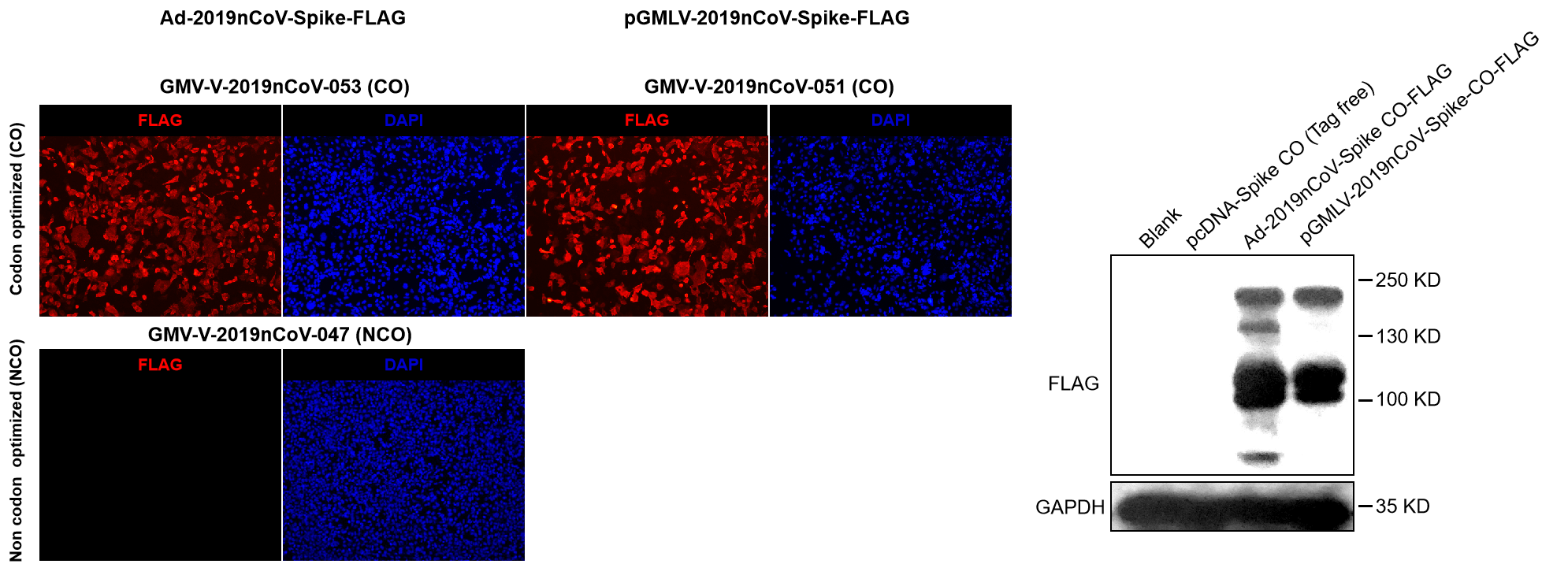
Figure. Validation of SARS-CoV-2 (2019nCoV) codon optimized (CO) Spike expression in HEK293 cell line by GeneMedi's adenoviral vector (GMV-V-2019nCoV-053, GMV-V-2019nCoV-047) and lentiviral vector (GMV-V-2019nCoV-051) with Immunofluorescence (left) and Western Blot (right) by FLAG antibody.
About SARS-CoV-2 (2019nCoV, novel Coronavirus) Spike protein, S1 Protein, Spike S1-NTD, Spike S1-CTD, Spike-RBD and Spike trimer protein.
1. SARS-CoV-2 (2019nCoV) Spike protein: SARS-CoV-2, a newly emerged pathogen spreading worldwide. The transmembrane spike (S) glycoprotein of SARS-CoV-2 that forms homotrimers protruding from the viral surface is known to mediate coronavirus entry into host cells. It has been reported that spike protein can bind with high affinity to human ACE2 and uses it as an entry receptor to invade target cells.
2. SARS-CoV-2 (2019nCoV) spike S1 protein: The spike protein is a large type I transmembrane glycoprotein comprises two functional subunits, S1 and S2. S1 subunit of spike protein is responsible for binding to the host cell receptor. S2 subunit is responsible for fusion of the viral and cellular membranes.
3. SARS-CoV-2 (2019nCoV) spike S1-NTD and S1-CTD: For most coronaviruses, the N-terminal domain (NTD) of the S1 subunit attaches to cellular carbohydrates and the C-terminal domain of S1 (S1-CTD) binds to a cellular protein receptor. Carbohydrate binding by the S1 N-terminal domain is thought to keep the virus in close proximity to the host cell surface, whereas engagement of specific protein receptors by the S1-CTD is thought to initiate a series of conformational changes in the spike that ultimately result in membrane fusion and delivery of the viral genome to the cytosol.
4. SARS-CoV-2 (2019nCoV) Spike RBD: S1 subunit of spike protein contains a receptor binding domain (RBD), which is responsible for recognizing the cell surface receptor.
5. SARS-CoV-2 (2019nCoV) Spike trimer protein: The transmembrane spike (S) glycoprotein of SARS-CoV-2 usually forms homotrimers protruding from the viral surface. S trimers are extensively decorated with N-linked glycans that are important for proper folding and for modulating accessibility to host proteases and neutralizing Abs. It has been found that S glycoprotein trimers in highly pathogenic human coronaviruses appear to exist in partially opened states, while they remain largely closed in human coronaviruses associated with common colds.
About SARS-CoV-2 (2019nCoV, novel Coronavirus) Envelop protein (Coronavirus E protein)
1. SARS-CoV-2 (2019nCoV) E protein: E protein is the smallest major structural proteins. It has a N-terminal ectodomain and a C-terminal endodomain with ion channel activity. During the replication cycle, E protein is abundantly expressed inside the infected cell, but only a small portion is incorporated into the virus envelope. The majority of the protein participates in viral assembly and budding. E protein is important in virus production and maturation. Recombinant CoVs without E have been shown to exhibit significantly reduced viral titres, crippled viral maturation, or yield incompetent progeny.
About SARS-CoV-2 (2019nCoV, novel Coronavirus) membrane protein (Coronavirus M protein)
1. SARS-CoV-2 (2019nCoV) M protein: Coronavirus M protein is believed to define the shape of the viral envelope,which contains three transmembrane domains. It has a small N-terminal glycosylated ectodomain and a much larger C-terminal endodomain that extends 6–8 nm into the viral particle. M protein usualy exists as a dimer, and may adopt two different conformations allowing it to promote membrane curvature as well as bind to the nucleocapsid.
About SARS-CoV-2 (2019nCoV, novel Coronavirus) Nucleocapsid protein (Coronavirus N protein)
1. SARS-CoV-2 (2019nCoV) N protein: Coronavirus N protein is required for coronavirus RNA synthesis, and has RNA chaperone activity that may be involved in template switch. Nucleocapsid protein is a most abundant protein of coronavirus. N protein packages the positive strand viral genome RNA into a helical ribonucleocapsid (RNP) and plays a fundamental role during virion assembly through its interactions with the viral genome and membrane protein M. Plays an important role in enhancing the efficiency of subgenomic viral RNA transcription as well as viral replication. Because of the conservation of N protein sequence and its strong immunogenicity, the N protein of coronavirus is chosen as a diagnostic tool.
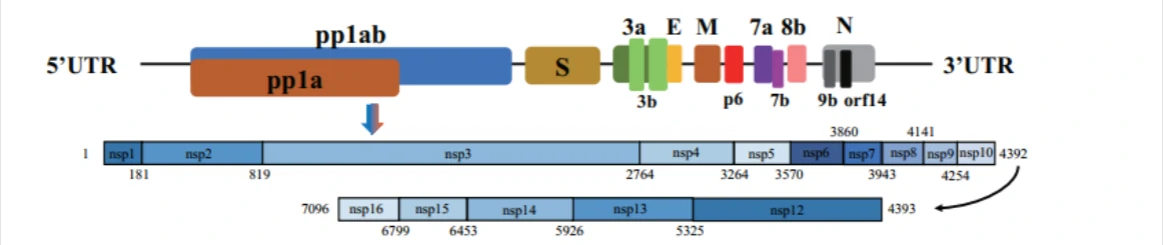
About SARS-CoV-2 (2019nCoV, novel Coronavirus)Non-structure protein (Nsp1-Nsp16)
2019nCoV contains 16 Non-structure protein (Nsp1-Nsp16) that may be drugable targets for antiviral compounds discovery against COVID-191.
| Non-structure proteins | Starting position (aa) | Ending position (aa) | Length (aa) |
| nsp1 | 1 | 180 | 180 |
| (leader protein) | |||
| nsp2 | 181 | 818 | 638 |
| nsp3 | 819 | 2763 | 1945 |
| (Papain-Like proteinase, PLpro) | |||
| nsp4 | 2764 | 3263 | 500 |
| nsp5 | 3264 | 3569 | 306 |
| (Mpro, Main proteinase, 3C-like proteinase) | |||
| nsp6 | 3570 | 3859 | 290 |
| nsp7 | 3860 | 3942 | 83 |
| nsp8 | 3943 | 4140 | 198 |
| nsp9 | 4141 | 4253 | 113 |
| nsp10 | 4254 | 4392 | 139 |
| (growth-factor-like protein) | |||
| nsp12 | 4393 | 5324 | 932 |
| (RdRP,NA-dependent RNA polymerase) | |||
| nsp13 | 5325 | 5925 | 601 |
| (RNA 5'-triphosphatase) | |||
| nsp14 | 5926 | 6452 | 527 |
| (3'-to-5' exonuclease) | |||
| nsp15 | 6453 | 6798 | 346 |
| (endoRNAse) | |||
| nsp16 | 6799 | 7096 | 298 |
| (2’O-MTase, 2'-O-ribose methyltransferase) |
1. Nsp3: Nsp3 (200 kDa) is the largest protein encoded by the coronavirus (CoV) genome. Nsp3 is an essential component of the replication and transcription complex. It comprises various domains, the organization of which differs between CoV genera, due to duplication or absence of some domains. However, the N-terminal region of the Nsp3 is highly conserved among CoV, containing a ubiquitin-like (Ubl) globular fold followed by a flexible, extended acidic-domain (AC domain) rich in glutamic acid (38%). Next to the AC domain is a catalytically active ADP-ribose-1"-phosphatase (ADRP, app-1"-pase) domain (also called macro domain or X domain) thought to play a role during synthesis of viral subgenomic RNAs. SARS Unique Domain (SUD), a domain not yet identified in other coronaviruses from alphacoronavirus and betacoronavirus, follows next. The SUD domain binds oligonucleotides known to form G-quadruplexes. Downstream of the SUD domain is a second Ubl domain and the catalytically active PLpro domain that proteolytically processes the Nsp1/2, Nsp2/3 and Nsp3/4 cleavage sites. Downstream of PLpro are found a nucleic acid-binding domain (NAB) with a nucleic acid chaperon function, which is conserved in betacoronavirus and gammacoronavirus, and one uncharacterized domain termed the marker domain (G2M). Following the G2M are two predicted double-pass transmembrane domains (TM1–2 and TM3–4), a putative metal binding region (ZN) and the Y domain of unknown function (subdomains Y1–3).
2. Nsp5: Nsp5 protease (3CLpro; Mpro) mediates processing at 11 distinct cleavage sites, including its own autoproteolysis, and is essential for virus replication. Nsp5 exhibits a conserved three-domain structure and catalytic residues.
3. Nsp10: Nsp10 (18 kDa) is well conserved among coronaviruses and encoded by ORF1a. It's thought to serve as an important multifunctional cofactor in replication. Nsp10 was shown to interact with itself, as well as with Nsp1, Nsp7, Nsp14, and Nsp16. The important role of Nsp10 is responsible for RNA synthesis. It was shown that a murine hepatitis virus (MHV) temperature-sensitive mutant carrying a non-synonymous mutation in the Nsp10 coding sequence had a defect in minus-strand RNA synthesis at non-permissive temperatures.
4. Nsp12: Nsp12 (102 kDa) is a multidomain RNA polymerase, which is the most conserved protein in coronaviruses. Nsp12 contains an RNA-dependent RNA polymerase (RdRp) domain in its C-terminal, which is essential for the viral replication and transcription.
5. Nsp16: Nsp16 is an SAM-dependent nucleoside-2’O-methyl-transferase (2’O-MTase). The mRNA cap for coronaviruses is completed by Nsp16, which ensures formation of a protective cap-1 structure that prevent recognition by either MDA5 or IFIT proteins. Finally, the NSP16/NSP10 complex finishes coronavirus capping process permitting viral infection with reduced host recognition.
About COVID-19 pandemic, Coronavirus (Coronavirus) and genome of SARS-CoV-2 (2019nCoV)
COVID-19 pandemic is caused by 2019nCoV (SARS-CoV-2, a novel coronavirus) infection.The 2019-nCoV genome was annotated to possess 14 ORFs encoding 27 proteins1.
| Gene name of 2019nCoV (SARS-CoV-2, a novel coronavirus) | Coding region(nt) | Protein length(aa) |
| orf1a | 266-13483 | 4405 |
| orf1ab | 266-13468, 13468-21555 | 7096 |
| S | 21563-25384 | 1273 |
| 3a | 25393-26220 | 275 |
| 3b | 25814-25882 | 22 |
| 26183-26281 | 32 | |
| E (envelope protei) | 26245-26472 | 75 |
| M (matrix protein) | 26523-27191 | 222 |
| p6 | 27202-27387 | 61 |
| 7a | 27394-27759 | 121 |
| 7b | 27756-27887 | 43 |
| 8b | 27894-28259 | 121 |
| 9b | 28284-28577 | 97 |
| N(nucleocapsid) | 28274-29533 | 419 |
| orf14 | 28734-28955 | 73 |
Collection of COVID-19 landscape knowledge base
Viral vector-based vaccine; DNA-based vaccine; RNA based vaccine
- A landscape for vaccine technology against infectious disease, COVID-19 and tumor.
COVID-19 landscape Knowledge Base
References
1. Wu, A. et al. Genome Composition and Divergence of the Novel Coronavirus (2019-nCoV) Originating in China. Cell Host Microbe, doi:10.1016/j.chom.2020.02.001 (2020).






