AAV engineering (AAV capsid evolution)
 AAV protocol Download
AAV protocol Download
AAV capsids evolution and tissue-specific AAV discovery & development service
The Genemedi help scientists and researchers in gene therapy vector development. Besides different scales of AAV production, we can also offer novel AAV discovery services with AAV re-engineering capsids library and sub-library generation based on different strategies, customized AAV capsids screening and tissue-specific AAV tropism validation.1. AAV engineering capsids library and sub-library generation
1) AAV-capsids peptides display library generation: 7 peptides & 14peptides, in unique site of different serotypes, basing on customized demand.
2) AAV-capsids DNA shuffling library generation;
3) Rational design AAV-capsids library ;
2. Customized AAV capsids screening in vitro and in vivo
1) Reengineered AAV assay strategy development;
2) Multi rounds of AAV re-engineered capsids in vitro and in vivo screening;
3) Candidates determination and analysis using Next generation sequencing (NGS) and single cell sequencing (SNS);
4) Optimization of AAV novel capsids candidates;
3. Tissue-specific AAV tropism validation in vitro and in vivo
In vitro: custom-demanded primary cells
In vivo: mouse, rat, Non-human primate (NHP)
Description
Although more than 100 AAV serotype variants have been found in nature, significant progress has been made for rAAV production, the transduction efficiency and specificity for some tissues and cell types in animals and primates is low. The non-targeted delivery by AAV requires high doses and potentially resulting in toxicities, problematic manufacturing burdens and suboptimal efficacy. Therefore, an important area in the development of AAV as a vector concerns the engineering of altered cell tropisms to increase rAAV mediated gene delivery efficiency and specificity. Among various AAV serotypes, the variations in transducing efficiencies could be caused by differences in their respective receptor contents on target cells. A great number of strategies for improving AAV delivery efficiency and specificity have been developed. Methodologically, the methods to improve AAV tropism and specificity can be divided into two categories. One is the assembly and recombination of proteins between different viruses, which can produce hybrid rAAVs. The other is to recombine, redesign, or introduce random mutations into the capsid protein of AAV by different methods to artificially increase the variance of AAV serotypes, and then screen the appropriate AAV serotypes. a) Hybrid rAAV Vectors The commonly used approach to generate hybrid rAAV vectors is transcapsidation, which is a process involving the packaging of the ITR from one AAV serotype into the capsid of another serotype, which may determine the tissue tropism of hybrids. For instance, if it is rAAV2/5 that is to be produced based on the helper-free system discussed above, the AAV packaging plasmid AAV-RC should be pRep2/Cap5, that is, the Rep gene is derived from AAV2, while the Capsid gene from AAV5. The packaging scheme is the same as the above-mentioned approach for AAV2 production. The resulting rAAV is named rAAV2/5, in which the genome is based on recombinant AAV2, while the capsid based on AAV5. The tissue tropism displayed by this rAAV2/5 hybrid virus is assumed to be the same with AAV5. b) AAV revolution There are two main ways to evolve AAV, one is rational design, the other is direct evolution (Figure4).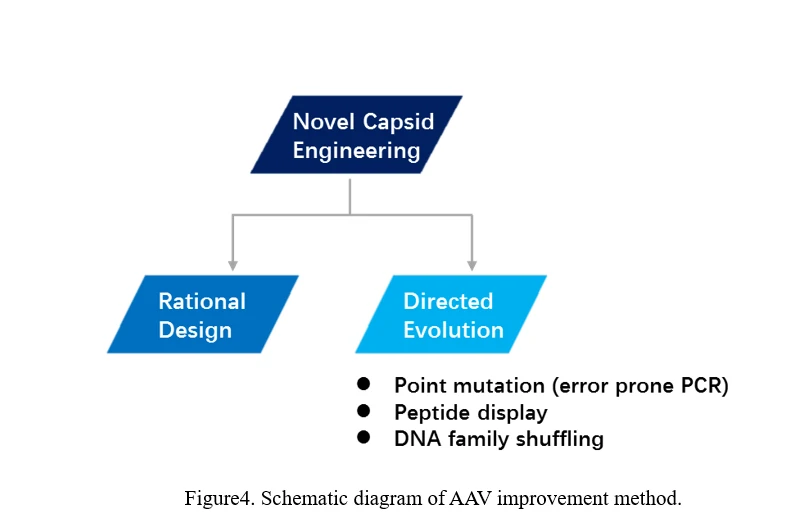
Rational Design
Some common methods of rational design AAV capsid, based on knowledge about delivery mechanisms coupled with AAV structural analysis [65], are listed as follows. (1) Phosphorylation of tyrosine residues in AAV capsid results in ubiquitylation and promotes proteasomal degradation of AAV virions, leading to the creation of vectors in which tyrosines were mutated to phenylalanines by site-directed mutagenesis [66]. (2) Analysis of potential docking sites for a murine IgG2a antibody with the AAV2 surface to determine sterically accessible candidate positions, which were then subjected to extensive site-directed mutagenesis to develop variants with reduced neutralization by antibodies in vitro [67]. (3) Incorporation of high-affinity ligands into the AAV capsid can confer binding to alternative cell surface receptors and thereby restrict or redirect viral tropism [68].AAV Directed Evolution
The conventional process of directed evolution is generating a library by error-prone PCR, peptide display or DNA family shuffling at firs. Then, this AAV viral library was packaged as a pool and purified, and selected under certain pressure in vitro or in vivo. After infection of the target cell or tissue, the viruses would recover from adenovirus-mediated replication or PCR amplification. Three or more rounds selection later, the highly infectious and specific AAV capsid species would be enriched (Figure 5) [69].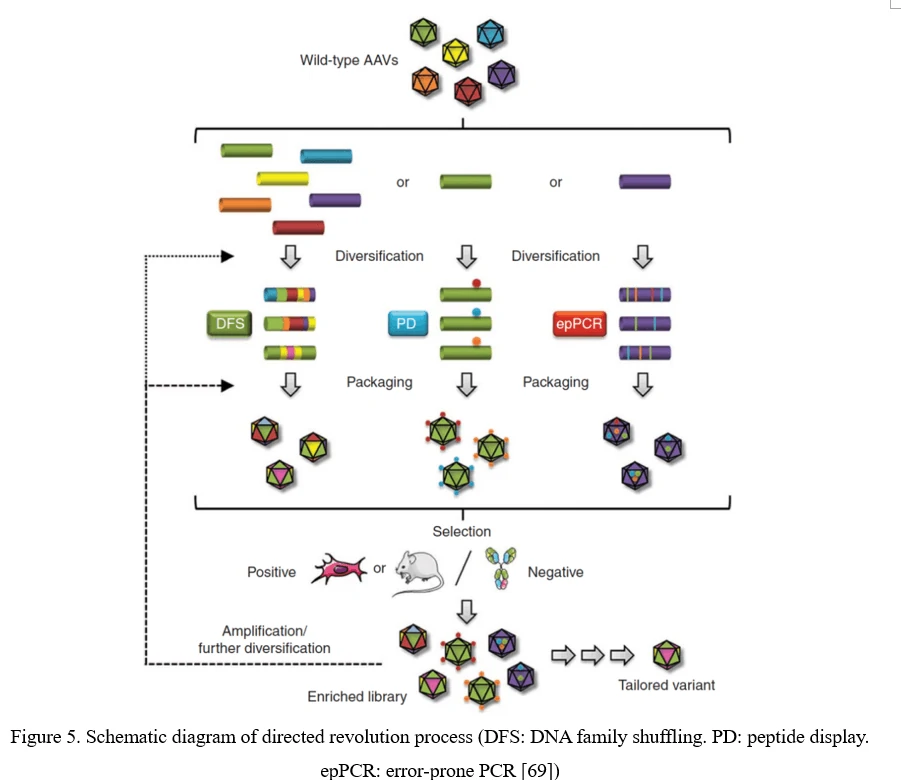
Point Mutation (Error-Prone PCR)
The simplest strategy for AAV diversification is mutagenesis of individual residues in the AAV viral capsid, either in a site-directed or in a randomized manner, expecting the introduced amino acid changes may alter essential AAV vector properties. Randomized mutations can be introduced to AAV capsid-encoding sequence by error-prone PCR (Figure 6). Although point mutations are easily introduced to AAV capsid-encoding sequence, there still exist problems. 1) Only 0.01–1% of AAV mutations are typically beneficial; 2) By far, number of clones to cover all combinations of multiple mutations within a single AAV capsid exceeds technical possibilities. It would be more useful to focus on a specific region of the AAV capsid, especially the surface-exposed areas and AAV capsid protrusions which are more likely to interact with cellular receptors and antibodies [70].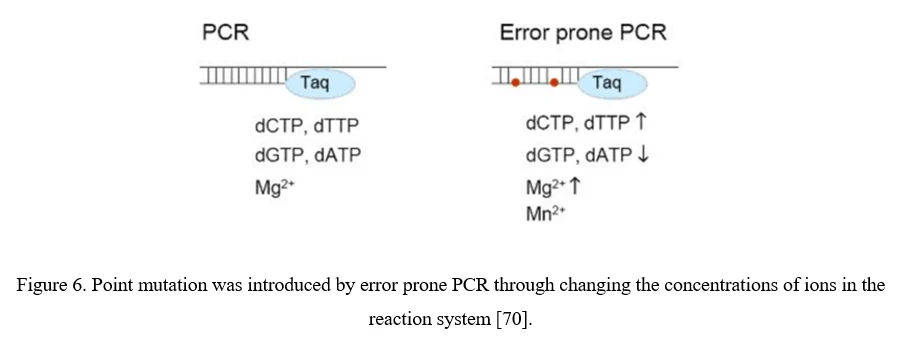
Peptide Display
Peptide insertion of novel receptor-binding elements on the AAV capsid surface can provide affinity for the receptor specifically expressed in the target cell type. To maximize success of this AAV peptide display strategy, it is critical to capitalize on sites in the AAV capsid that mediate surface exposure of the peptide. In the case of AAV2, it can be achieved by insertion before or after arginine 588 (R588), one of the two vital residues required for AAV2 binding to HSPG. In the AAV2-7m8 variant, 7m8 disrupts the HSPG-binding motif, taking the arginines apart (Figure 7) [71].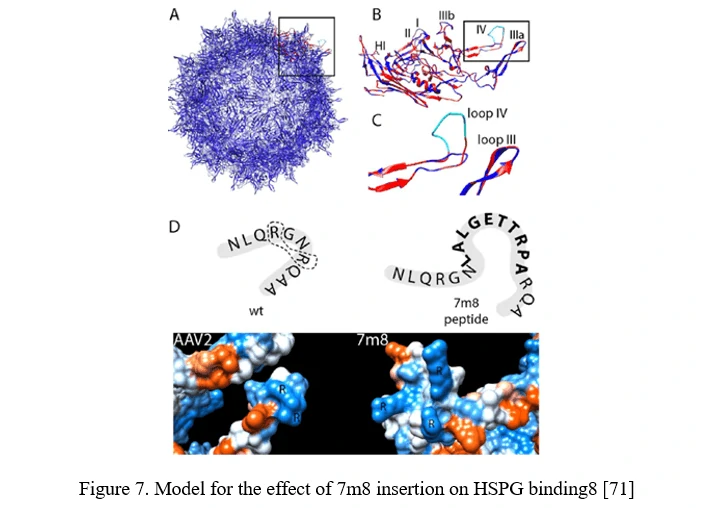
AAV family shuffling (DNA Shuffling of AAV capsids)
A chimeric capsid–encoding library can be generated by AAV family shuffling (Figure 8). Fragments of various AAV capsid genes are re-assembled based on partial homologies (typically >80% for most AAV serotypes). This library contains chimeras with broad diversity in cell tropism and neutralizing antibody resistance. This approach can also harness and blend functions from newly identified AAV serotypes, or even rationally designed AAV capsid mutants, to potentially create a combinatorially limitless number of novel variants with new properties [72].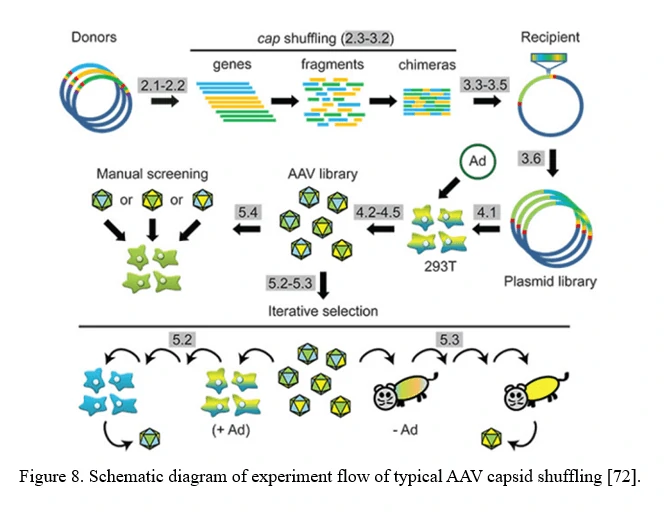
View Adeno-associated virus (AAV vector) Knowledge Base>>
Other knowledge bases
AAV Knowledge Base CRISPR/Cas9-gRNA system Knowledge Base |
Adenovirus Knowledge Base Transfection Knowledge Base |
Lentivirus Knowledge Base Advanced cell therapy |
AAV(Adeno-Associated Virus) vector system
AAV Rep-Cap plasmids (serotypes-specific AAV RC plasmids)




